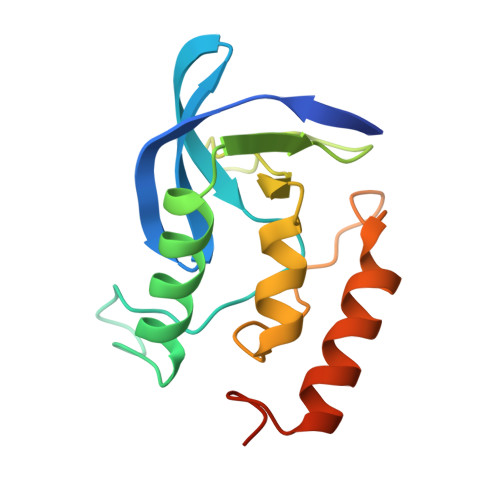
Crystal structures of the binary Ca2+ and pdTp complexes and the ternary complex of the Asp21-->Glu mutant of staphylococcal nuclease. Implications for catalysis and ligand binding.
Libson, A.M., Gittis, A.G., Lattman, E.E.(1994) Biochemistry 33: 8007-8016
- PubMed: 8025105
- Primary Citation of Related Structures:
1ENA, 1ENC, 2ENB - PubMed Abstract:
The crystal structure of the Asp21-->Glu mutant (D21E) of staphylococcal nuclease (SNase) has been determined in three different complex forms. The structure of the D21E ternary complex in which D21E is bound to both Ca2+ and the transition-state analogue, thymidine 3',5'-diphosphate (pdTp), was determined to 1.95-A resolution. The structures of both binary complexes, D21E bound either to Ca2+ or pdTp, were determined to 2.15- and 2.05-A resolution, respectively. In the ternary structure, we find a 1.5-A movement of the Ca2+ in the active site, evidence of bidentate coordination of Ca2+ by Glu21 and inner-sphere coordination of the Ca2+ by Glu43. Comparison of the D21E binary structures with the ternary model shows large movements of active site side chains expected to play a direct role in catalysis. Glu43 moves in the binary nucleotide complex, whereas Arg35 is oriented differently in the binary metal complex. From these changes, we seek to explain the basis for the 1500-fold decrease in Vmax of D21E relative to wild-type SNase (WT). Furthermore, we describe direct structural evidence which explains the cooperativity of Ca2+ and pdTp binding in the ternary complex relative to that of the binary complexes.
Organizational Affiliation:
Department of Biophysics & Biophysical Chemistry, Johns Hopkins School of Medicine, Baltimore, Maryland 21205.