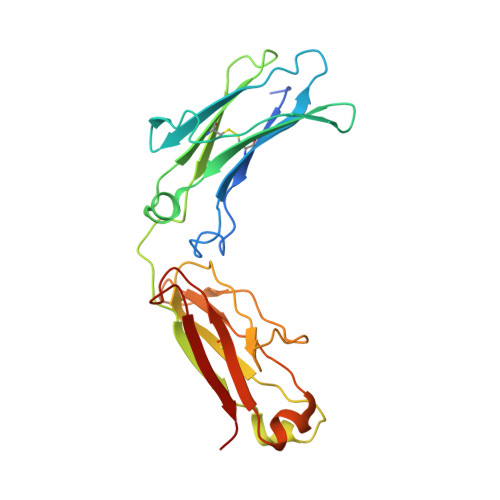
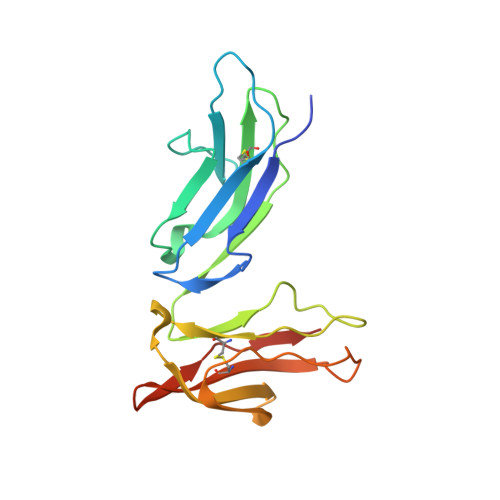
The structure of a human type III Fcgamma receptor in complex with Fc
Radaev, S., Motyka, S., Fridman, W.-H., Sautes-Fridman, C., Sun, P.D.(2001) J Biol Chem 276: 16469-16477
- PubMed: 11297532
- DOI: https://doi.org/10.1074/jbc.M100350200
- Primary Citation of Related Structures:
1T83, 1T89 - PubMed Abstract:
Fcgamma receptors mediate antibody-dependent inflammatory responses and cytotoxicity as well as certain autoimmune dysfunctions. Here we report the crystal structure of a human Fc receptor (FcgammaRIIIB) in complex with an Fc fragment of human IgG1 determined from orthorhombic and hexagonal crystal forms at 3.0- and 3.5-A resolution, respectively. The refined structures from the two crystal forms are nearly identical with no significant discrepancies between the coordinates. Regions of the C-terminal domain of FcgammaRIII, including the BC, C'E, FG loops, and the C' beta-strand, bind asymmetrically to the lower hinge region, residues Leu(234)-Pro(238), of both Fc chains creating a 1:1 receptor-ligand stoichiometry. Minor conformational changes are observed in both the receptor and Fc upon complex formation. Hydrophobic residues, hydrogen bonds, and salt bridges are distributed throughout the receptor.Fc interface. Sequence comparisons of the receptor-ligand interface residues suggest a conserved binding mode common to all members of immunoglobulin-like Fc receptors. Structural comparison between FcgammaRIII.Fc and FcepsilonRI.Fc complexes highlights the differences in ligand recognition between the high and low affinity receptors. Although not in direct contact with the receptor, the carbohydrate attached to the conserved glycosylation residue Asn(297) on Fc may stabilize the conformation of the receptor-binding epitope on Fc. An antibody-FcgammaRIII model suggests two possible ligand-induced receptor aggregations.
Organizational Affiliation:
Structural Biology Section, Laboratory of Immunogenetics, NIAID, National Institutes of Health, Rockville, Maryland 20852, USA.