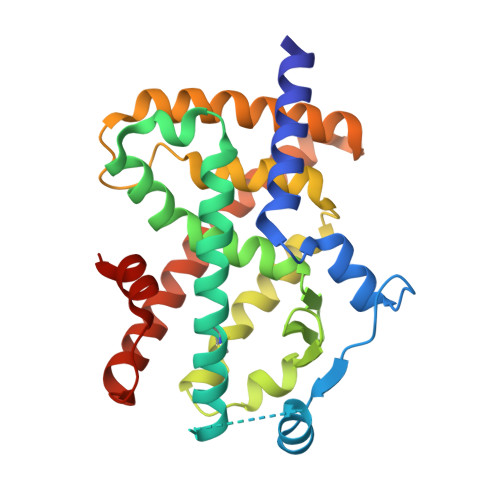
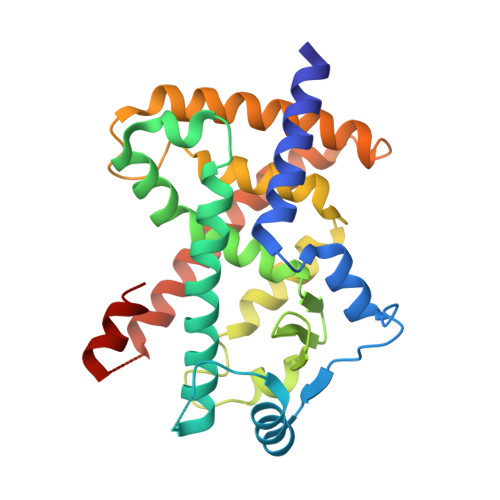
Partial Agonists Activate PPARgamma Using a Helix 12 Independent Mechanism
Bruning, J.B., Chalmers, M.J., Prasad, S., Busby, S.A., Kamenecka, T.M., He, Y., Nettles, K.W., Griffin, P.R.(2007) Structure 15: 1258-1271
- PubMed: 17937915
- DOI: https://doi.org/10.1016/j.str.2007.07.014
- Primary Citation of Related Structures:
2Q59, 2Q5P, 2Q5S, 2Q61, 2Q6R, 2Q6S - PubMed Abstract:
Binding to helix 12 of the ligand-binding domain of PPARgamma is required for full agonist activity. Previously, the degree of stabilization of the activation function 2 (AF-2) surface was thought to correlate with the degree of agonism and transactivation. To examine this mechanism, we probed structural dynamics of PPARgamma with agonists that induced graded transcriptional responses. Here we present crystal structures and amide H/D exchange (HDX) kinetics for six of these complexes. Amide HDX revealed each ligand induced unique changes to the dynamics of the ligand-binding domain (LBD). Full agonists stabilized helix 12, whereas intermediate and partial agonists did not at all, and rather differentially stabilized other regions of the binding pocket. The gradient of PPARgamma transactivation cannot be accounted for solely through changes to the dynamics of AF-2. Thus, our understanding of allosteric signaling must be extended beyond the idea of a dynamic helix 12 acting as a molecular switch.
Organizational Affiliation:
Department of Cancer Biology, The Scripps Research Institute, 5353 Parkside Drive, Jupiter, FL 33458, USA.