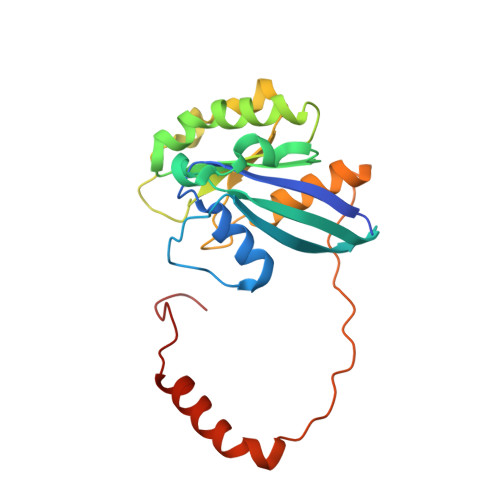
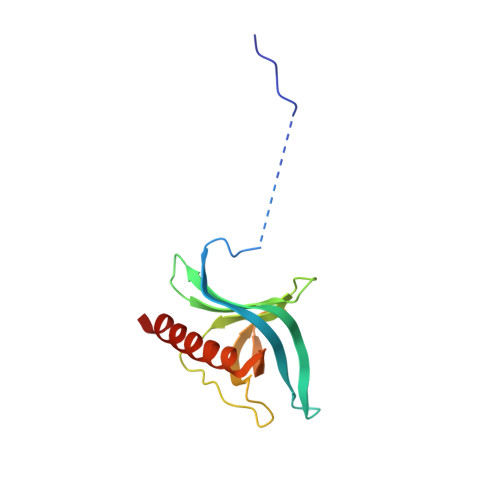
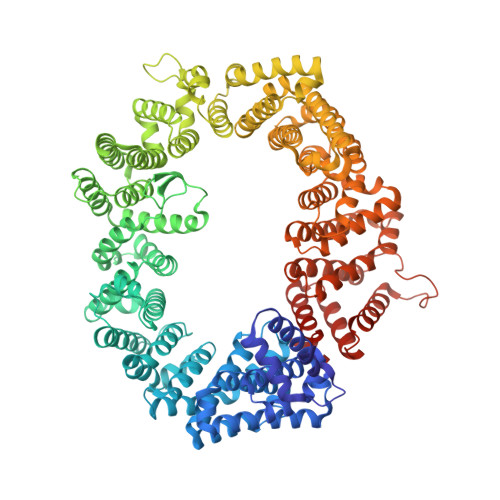
Nuclear export inhibitors avert progression in preclinical models of inflammatory demyelination.
Haines, J.D., Herbin, O., de la Hera, B., Vidaurre, O.G., Moy, G.A., Sun, Q., Fung, H.Y., Albrecht, S., Alexandropoulos, K., McCauley, D., Chook, Y.M., Kuhlmann, T., Kidd, G.J., Shacham, S., Casaccia, P.(2015) Nat Neurosci 18: 511-520
- PubMed: 25706475
- DOI: https://doi.org/10.1038/nn.3953
- Primary Citation of Related Structures:
4WVF - PubMed Abstract:
Axonal damage has been associated with aberrant protein trafficking. We examined a newly characterized class of compounds that target nucleo-cytoplasmic shuttling by binding to the catalytic groove of the nuclear export protein XPO1 (also known as CRM1, chromosome region maintenance protein 1). Oral administration of reversible CRM1 inhibitors in preclinical murine models of demyelination significantly attenuated disease progression, even when started after the onset of paralysis. Clinical efficacy was associated with decreased proliferation of immune cells, characterized by nuclear accumulation of cell cycle inhibitors, and preservation of cytoskeletal integrity even in demyelinated axons. Neuroprotection was not limited to models of demyelination, but was also observed in another mouse model of axonal damage (that is, kainic acid injection) and detected in cultured neurons after knockdown of Xpo1, the gene encoding CRM1. A proteomic screen for target molecules revealed that CRM1 inhibitors in neurons prevented nuclear export of molecules associated with axonal damage while retaining transcription factors modulating neuroprotection.
Organizational Affiliation:
Department of Neuroscience, Icahn School of Medicine at Mount Sinai, New York, New York, USA.