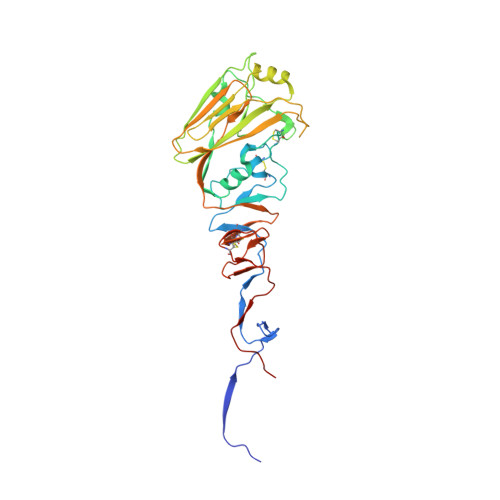
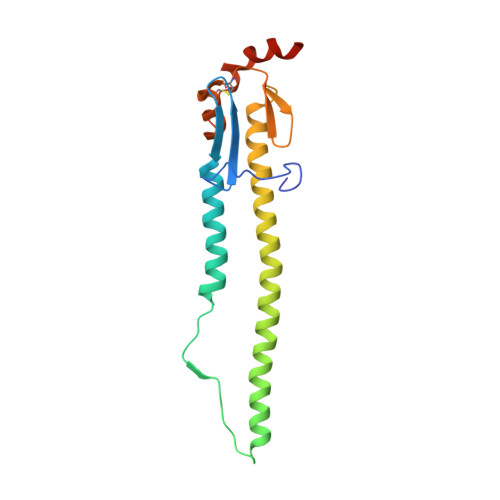
Conserved epitope on influenza-virus hemagglutinin head defined by a vaccine-induced antibody.
Raymond, D.D., Bajic, G., Ferdman, J., Suphaphiphat, P., Settembre, E.C., Moody, M.A., Schmidt, A.G., Harrison, S.C.(2018) Proc Natl Acad Sci U S A 115: 168-173
- PubMed: 29255041
- DOI: https://doi.org/10.1073/pnas.1715471115
- Primary Citation of Related Structures:
5W6C, 5W6G - PubMed Abstract:
Circulating influenza viruses evade neutralization in their human hosts by acquiring escape mutations at epitopes of prevalent antibodies. A goal for next-generation influenza vaccines is to reduce escape likelihood by selectively eliciting antibodies recognizing conserved surfaces on the viral hemagglutinin (HA). The receptor-binding site (RBS) on the HA "head" and a region near the fusion peptide on the HA "stem" are two such sites. We describe here a human antibody clonal lineage, designated CL6649, members of which bind a third conserved site ("lateral patch") on the side of the H1-subtype, HA head. A crystal structure of HA with bound Fab6649 shows the conserved antibody footprint. The site was invariant in isolates from 1977 (seasonal) to 2012 (pdm2009); antibodies in CL6649 recognize HAs from the entire period. In 2013, human H1 viruses acquired mutations in this epitope that were retained in subsequent seasons, prompting modification of the H1 vaccine component in 2017. The mutations inhibit Fab6649 binding. We infer from the rapid spread of these mutations in circulating H1 influenza viruses that the previously subdominant, conserved lateral patch had become immunodominant for individuals with B-cell memory imprinted by earlier H1 exposure. We suggest that introduction of the pdm2009 H1 virus, to which most of the broadly prevalent, neutralizing antibodies did not bind, conferred a selective advantage in the immune systems of infected hosts to recall of memory B cells that recognized the lateral patch, the principal exposed epitope that did not change when pdm2009 displaced previous seasonal H1 viruses.
Organizational Affiliation:
Laboratory of Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, MA 02115.