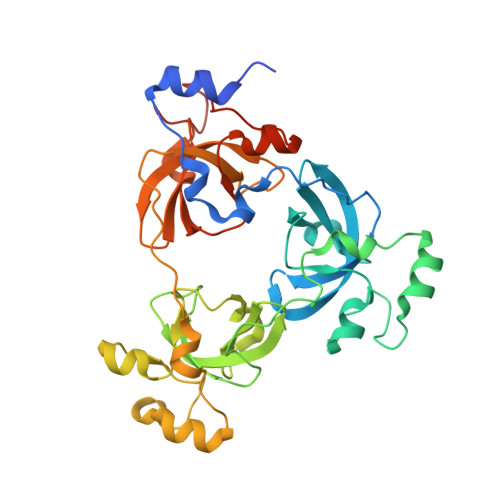
Structural Basis for Lower Lysine Methylation State-Specific Readout by MBT Repeats of L3MBTL1 and an Engineered PHD Finger.
Li, H., Fischle, W., Wang, W., Duncan, E.M., Liang, L., Murakami-Ishibe, S., Allis, C.D., Patel, D.J.(2007) Mol Cell 28: 677-691
- PubMed: 18042461
- DOI: https://doi.org/10.1016/j.molcel.2007.10.023
- Primary Citation of Related Structures:
2RHI, 2RHU, 2RHX, 2RHY, 2RHZ, 2RI2, 2RI3, 2RI5, 2RI7 - PubMed Abstract:
Human L3MBTL1, which contains three malignant brain tumor (MBT) repeats, binds monomethylated and dimethylated lysines, but not trimethylated lysines, in several histone sequence contexts. In crystal structures of L3MBTL1 complexes, the monomethyl- and dimethyllysines insert into a narrow and deep cavity of aromatic residue-lined pocket 2, while a proline ring inserts into shallower pocket 1. We have also engineered a single Y to E substitution within the aromatic cage of the BPTF PHD finger, resulting in a reversal of binding preference from trimethyl- to dimethyllysine in an H3K4 sequence context. In both the "cavity insertion" (L3MBTL1) and "surface groove" (PHD finger) modes of methyllysine recognition, a carboxylate group both hydrogen bonds and ion pairs to the methylammonium proton. Our structural and binding studies of these two modules provide insights into the molecular principles governing the decoding of lysine methylation states, thereby highlighting a methylation state-specific layer of histone mark readout impacting on epigenetic regulation.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10021, USA.