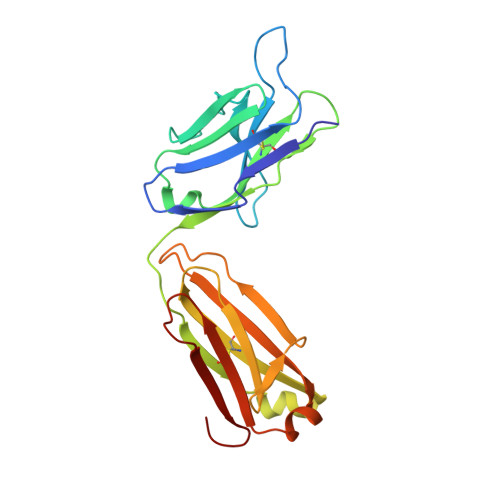
A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV.
Yuan, M., Wu, N.C., Zhu, X., Lee, C.D., So, R.T.Y., Lv, H., Mok, C.K.P., Wilson, I.A.(2020) Science 368: 630-633
- PubMed: 32245784
- DOI: https://doi.org/10.1126/science.abb7269
- Primary Citation of Related Structures:
6W41 - PubMed Abstract:
The outbreak of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome-coronavirus 2 (SARS-CoV-2) has now become a pandemic, but there is currently very little understanding of the antigenicity of the virus. We therefore determined the crystal structure of CR3022, a neutralizing antibody previously isolated from a convalescent SARS patient, in complex with the receptor binding domain (RBD) of the SARS-CoV-2 spike (S) protein at 3.1-angstrom resolution. CR3022 targets a highly conserved epitope, distal from the receptor binding site, that enables cross-reactive binding between SARS-CoV-2 and SARS-CoV. Structural modeling further demonstrates that the binding epitope can only be accessed by CR3022 when at least two RBDs on the trimeric S protein are in the "up" conformation and slightly rotated. These results provide molecular insights into antibody recognition of SARS-CoV-2.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.