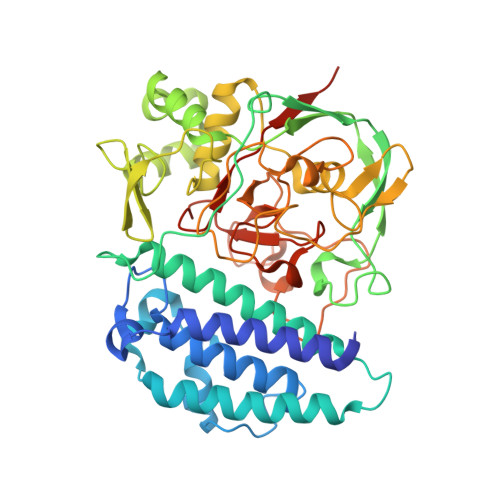
Structure of the Sulfoxide Synthase EgtB from the Ergothioneine Biosynthetic Pathway.
Goncharenko, K.V., Vit, A., Blankenfeldt, W., Seebeck, F.P.(2015) Angew Chem Int Ed Engl 54: 2821-2824
- PubMed: 25597398
- DOI: https://doi.org/10.1002/anie.201410045
- Primary Citation of Related Structures:
4X8B, 4X8D, 4X8E - PubMed Abstract:
The non-heme iron enzyme EgtB catalyzes O2 -dependent C-S bond formation between γ-glutamyl cysteine and N-α-trimethyl histidine as the central step in ergothioneine biosynthesis. Both, the catalytic activity and the architecture of EgtB are distinct from known sulfur transferases or thiol dioxygenases. The crystal structure of EgtB from Mycobacterium thermoresistibile in complex with γ-glutamyl cysteine and N-α-trimethyl histidine reveals that the two substrates and three histidine residues serve as ligands in an octahedral iron binding site. This active site geometry is consistent with a catalytic mechanism in which C-S bond formation is initiated by an iron(III)-complexed thiyl radical attacking the imidazole ring of N-α-trimethyl histidine.
Organizational Affiliation:
Department for Chemistry, University of Basel, St. Johanns-Ring 19, 4056 Basel (Switzerland).