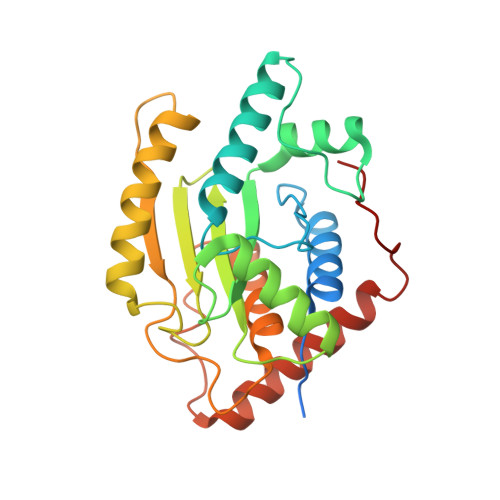
Structural Basis of Substrate Recognition and Covalent Inhibition of Cdu1 from Chlamydia trachomatis.
Ramirez, Y.A., Adler, T.B., Altmann, E., Klemm, T., Tiesmeyer, C., Sauer, F., Kathman, S.G., Statsyuk, A.V., Sotriffer, C., Kisker, C.(2018) ChemMedChem 13: 2014-2023
- PubMed: 30028574
- DOI: https://doi.org/10.1002/cmdc.201800364
- Primary Citation of Related Structures:
6FDK, 6FDQ, 6FDU - PubMed Abstract:
Based on the similarity between the active sites of the deubiquitylating and deneddylating enzyme ChlaDub1 (Cdu1) and the evolutionarily related protease adenain, a target-hopping screening approach on a focused set of adenain inhibitors was investigated. The cyanopyrimidine-based inhibitors identified represent the first active-site-directed small-molecule inhibitors of Cdu1. High-resolution crystal structures of Cdu1 in complex with two covalently bound cyanopyrimidines, as well as with its substrate ubiquitin, were obtained. These structural data were complemented by enzymatic assays and covalent docking studies to provide insight into the substrate recognition of Cdu1, active-site pocket flexibility and potential hotspots for ligand interaction. Combined, these data provide a strong basis for future structure-guided medicinal chemistry optimization of this cyanopyrimidine scaffold into more potent and selective Cdu1 inhibitors.
Organizational Affiliation:
Rudolf Virchow Center for Experimental Biomedicine, Institute of Structural Biology, University of Würzburg, Josef-Schneider-Straße 2, 97080, Würzburg, Germany.