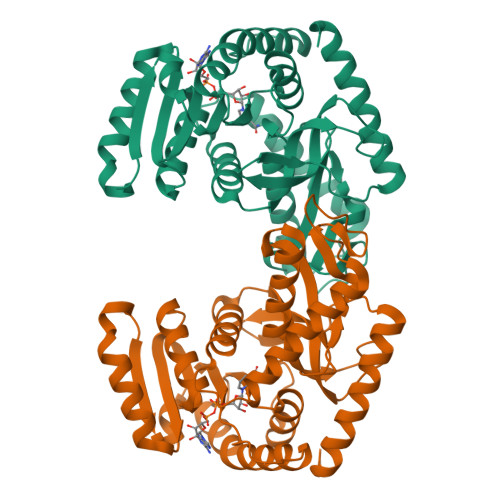
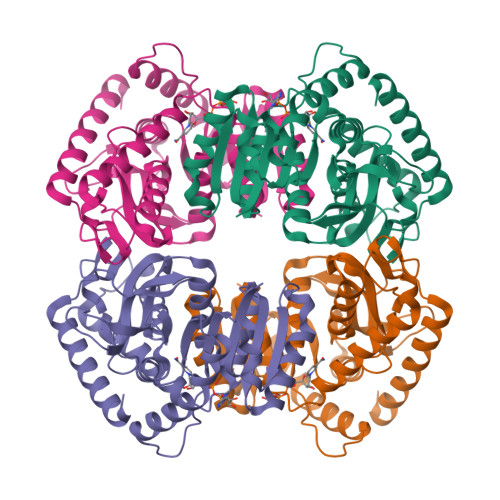
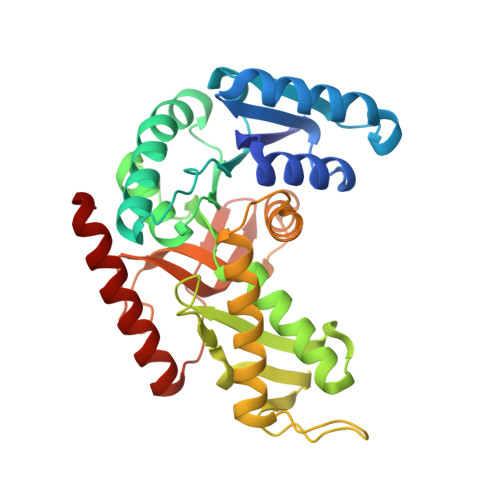
Molecular basis of allosteric activation of bacterial L-lactate dehydrogenase.
Iwata, S., Ohta, T.(1993) J Mol Biology 230: 21-27
- PubMed: 8450537
- DOI: https://doi.org/10.1006/jmbi.1993.1122
- Primary Citation of Related Structures:
1LLD - PubMed Abstract:
The three-dimensional structure of allosteric L-lactate dehydrogenase from Bifidobacterium longum, the first example of a T-state structure of L-lactate dehydrogenase, has been determined to 2.0 A. A comparative study of this structure with the previously reported R-state structure from Bacillus stearothermophilus has revealed the allosteric activation mechanism of the bacterial L-lactate dehydrogenase. The fructose 1,6-bisphosphate-induced conformational change at the effector site and the substrate affinity change at the activity site are clearly shown at a molecular level. Coupling of these changes can be simply explained by a set of concerted rotations between subunits in the tetramer of the enzyme. This T to R transition is the first example for a tetrameric allosteric protein where the rotations occur around each of three axes of symmetry.
Organizational Affiliation:
Photon Factory, National Laboratory for High Energy Physics, Ibaraki-ken, Japan.