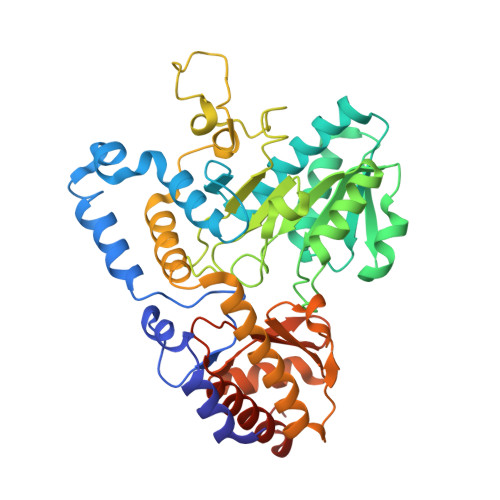
Three-Dimensional Structure of Kynureninase from Pseudomonas fluorescens.
Momany, C., Levdikov, V., Blagova, L., Lima, S., Phillips, R.S.(2004) Biochemistry 43: 1193-1203
- PubMed: 14756555
- DOI: https://doi.org/10.1021/bi035744e
- Primary Citation of Related Structures:
1QZ9 - PubMed Abstract:
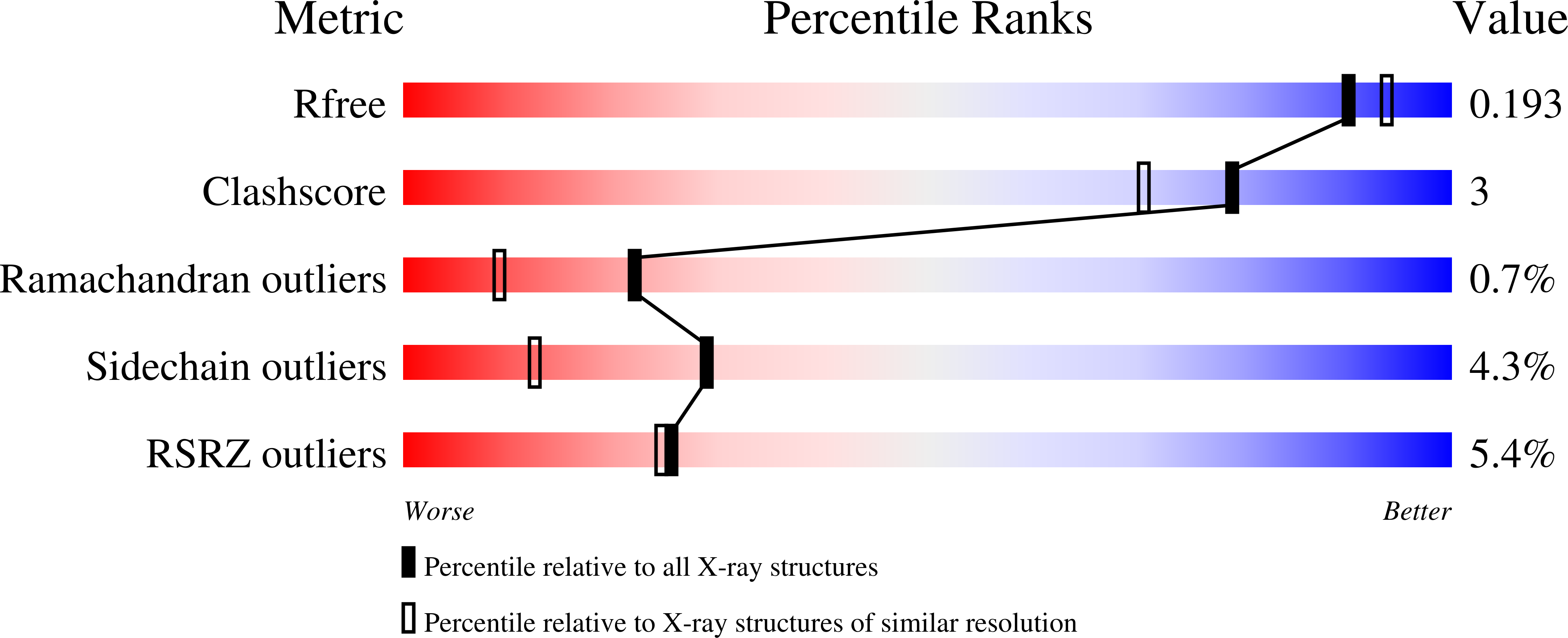
Kynureninase [E.C. 3.7.1.3] is a pyridoxal-5'-phosphate (PLP)-dependent enzyme that catalyzes the hydrolytic cleavage of l-kynurenine to anthranilic acid and l-alanine. Sequence alignment with other PLP-dependent enzymes indicated that kynureninase is in subgroup IVa of the aminotransferases, along with nifS, CsdB, and serine-pyruvate aminotransferase, which suggests that kynureninase has an aminotransferase fold. Crystals of Pseudomonas fluorescens kynureninase were obtained, and the structure was solved by molecular replacement using the CsdB coordinates combined with multiple isomorphous heavy atom replacement. The coordinates were deposited in the PDB (ID code 1QZ9). The structure, refined to an R factor of 15.5% to 1.85 A resolution, is dimeric and has the aminotransferase fold. The structure also confirms the prediction from sequence alignment that Lys-227 is the PLP-binding residue in P. fluorescens kynureninase. The conserved Asp-201, expected for an aminotransferase fold, is located near the PLP nitrogen, but Asp-132 is also strictly conserved and at a similar distance from the pyridinium nitrogen. Mutagenesis of both conserved aspartic acids shows that both contribute equally to PLP binding, but Asp-201 has a greater role in catalysis. The structure shows that Tyr-226 donates a hydrogen bond to the phosphate of PLP. Unusual among PLP-dependent enzymes, Trp-256, which is also strictly conserved in kynureninases from bacteria to humans, donates a hydrogen bond to the phosphate through the indole N1-hydrogen.
Organizational Affiliation:
Department of Chemistry, University of Georgia, Athens, Georgia 30602, USA.