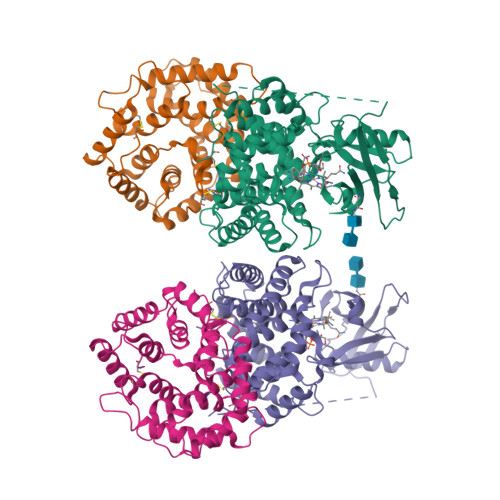
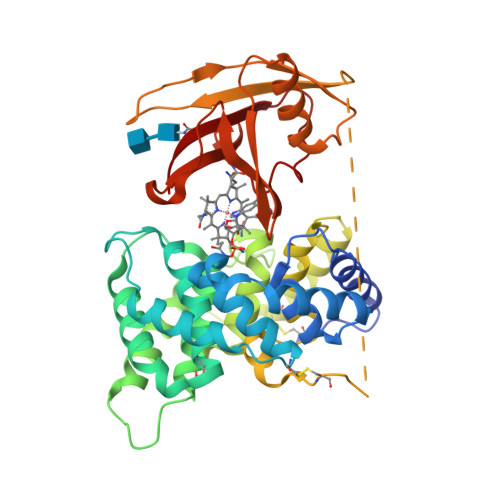
Crystal structure of human intrinsic factor: Cobalamin complex at 2.6-A resolution
Mathews, F.S., Gordon, M.M., Chen, Z., Rajashankar, K.R., Ealick, S.E., Alpers, D.H., Sukumar, N.(2007) Proc Natl Acad Sci U S A 104: 17311-17316
- PubMed: 17954916
- DOI: https://doi.org/10.1073/pnas.0703228104
- Primary Citation of Related Structures:
2PMV - PubMed Abstract:
The structure of intrinsic factor (IF) in complex with cobalamin (Cbl) was determined at 2.6-A resolution. The overall fold of the molecule is that of an alpha(6)/alpha(6) barrel. It is a two-domain protein, and the Cbl is bound at the interface of the domains in a base-on conformation. Surprisingly, two full-length molecules, each comprising an alpha- and a beta-domain and one Cbl, and two truncated molecules with only an alpha- domain are present in the same asymmetric unit. The environment around Cbl is dominated by uncharged residues, and the sixth coordinate position of Co(2+) is empty. A detailed comparison between the IF-B12 complex and another Cbl transport protein complex, trans-Cbl-B12, has been made. The pH effect on the binding of Cbl analogues in transport proteins is analyzed. A possible basis for the lack of interchangeability of human and rat IF receptors is presented.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110, USA.