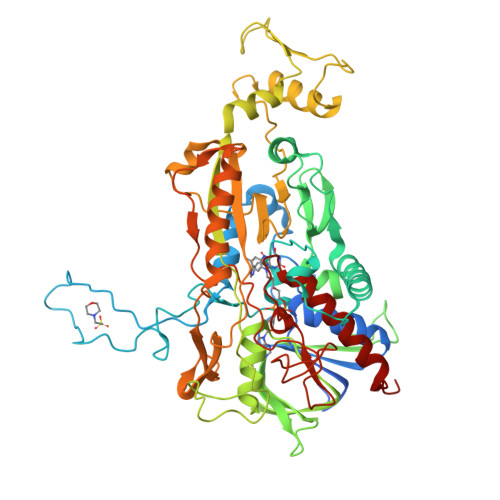
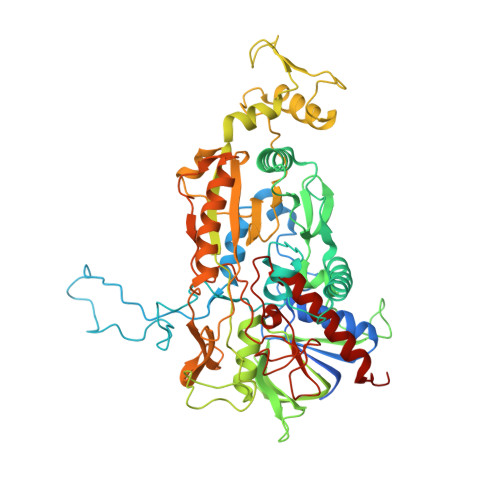
A thermostable triple mutant of pyranose 2-oxidase from Trametes multicolor with improved properties for biotechnological applications
Spadiut, O., Radakovits, K., Pisanelli, I., Salaheddin, C., Yamabhai, M., Tan, T.-C., Divne, C., Haltrich, D.(2009) Biotechnol J 4: 525-534
- PubMed: 19291706
- DOI: https://doi.org/10.1002/biot.200800260
- Primary Citation of Related Structures:
3FDY - PubMed Abstract:
In order to increase the thermal stability and the catalytic properties of pyranose oxidase (P2Ox) from Trametes multicolor toward its poor substrate D-galactose and the alternative electron acceptor 1,4-benzoquinone (1,4-BQ), we designed the triple-mutant T169G/E542K/V546C. Whereas the wild-type enzyme clearly favors D-glucose as its substrate over D-galactose [substrate selectivity (k(cat)/K(M))(Glc)/(k(cat)/K(M))(Gal) = 172], the variant oxidizes both sugars equally well [(k(cat)/K(M))(Glc)/(k(cat)/K(M))(Gal) = 0.69], which is of interest for food biotechnology. Furthermore, the variant showed lower K(M) values and approximately ten-fold higher k(cat) values for 1,4-BQ when D-galactose was used as the saturating sugar substrate, which makes this enzyme particularly attractive for use in biofuel cells and enzyme-based biosensors. In addition to the altered substrate specificity and reactivity, this mutant also shows significantly improved thermal stability. The half life time at 60 degrees C was approximately 10 h, compared to 7.6 min for the wild-type enzyme. We performed successfully small-scale bioreactor pilot conversion experiments of D-glucose/D-galactose mixtures at both 30 and 50 degrees C, showing the usefulness of this P2Ox variant in biocatalysis as well as the enhanced thermal stability of the enzyme. Moreover, we determined the crystal structure of the mutant in its unligated form at 1.55 A resolution. Modeling D-galactose in position for oxidation at C2 into the mutant active site shows that substituting Thr for Gly at position 169 favorably accommodates the axial C4 hydroxyl group that would otherwise clash with Thr169 in the wild-type.
Organizational Affiliation:
Department of Food Sciences and Technology, BOKU, Vienna, Austria.