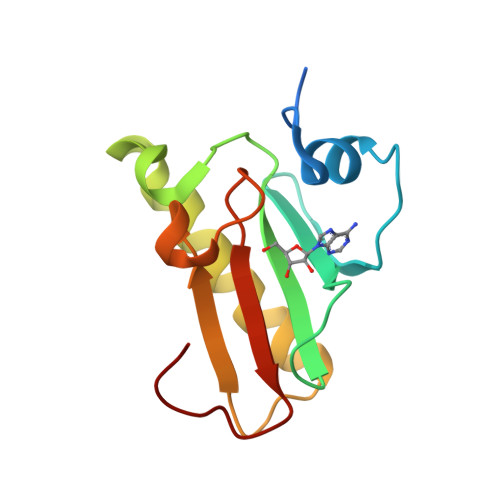
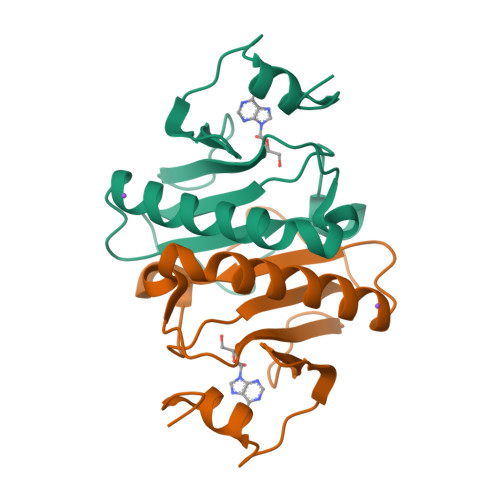
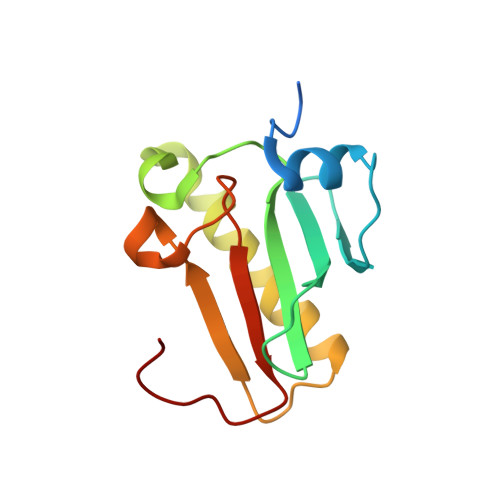
Histidine Triad Nucleotide-binding Protein 1 (HINT-1) Phosphoramidase Transforms Nucleoside 5'-O-Phosphorothioates to Nucleoside 5'-O-Phosphates.
Ozga, M., Dolot, R., Janicka, M., Kaczmarek, R., Krakowiak, A.(2010) J Biological Chem 285: 40809-40818
- PubMed: 20940308
- DOI: https://doi.org/10.1074/jbc.M110.162065
- Primary Citation of Related Structures:
3O1C, 3O1X, 3O1Z - PubMed Abstract:
Nucleoside 5'-O-phosphorothioates are formed in vivo as primary products of hydrolysis of oligo(nucleoside phosphorothioate)s (PS-oligos) that are applied as antisense therapeutic molecules. The biodistribution of PS-oligos and their pharmacokinetics have been widely reported, but little is known about their subsequent decay inside the organism. We suggest that the enzyme responsible for nucleoside 5'-O-monophosphorothioate ((d)NMPS) metabolism could be histidine triad nucleotide-binding protein 1 (Hint-1), a phosphoramidase belonging to the histidine triad (HIT) superfamily that is present in all forms of life. An additional, but usually ignored, activity of Hint-1 is its ability to catalyze the conversion of adenosine 5'-O-monophosphorothioate (AMPS) to 5'-O-monophosphate (AMP). By mutagenetic and biochemical studies, we defined the active site of Hint-1 and the kinetic parameters of the desulfuration reaction (P-S bond cleavage). Additionally, crystallographic analysis (resolution from 1.08 to 1.37 Å) of three engineered cysteine mutants showed the high similarity of their structures, which were not very different from the structure of WT Hint-1. Moreover, we found that not only AMPS but also other ribonucleoside and 2'-deoxyribonucleoside phosphorothioates are desulfurated by Hint-1 at the following relative rates: GMPS > AMPS > dGMPS ≥ CMPS > UMPS > dAMPS ≫ dCMPS > TMPS, and during the reaction, hydrogen sulfide, which is thought to be the third gaseous mediator, was released.
Organizational Affiliation:
Department of Bioorganic Chemistry, Centre of Molecular and Macromolecular Studies, Polish Academy of Sciences, Lodz 90-363, Poland.