Cyclic GMP-AMP Synthase Is Activated by Double-Stranded DNA-Induced Oligomerization.
Li, X., Shu, C., Yi, G., Chaton, C.T., Shelton, C.L., Diao, J., Zuo, X., Kao, C.C., Herr, A.B., Li, P.(2013) Immunity 39: 1019-1031
- PubMed: 24332030
- DOI: https://doi.org/10.1016/j.immuni.2013.10.019
- Primary Citation of Related Structures:
4LEV, 4LEW, 4LEY, 4LEZ - PubMed Abstract:
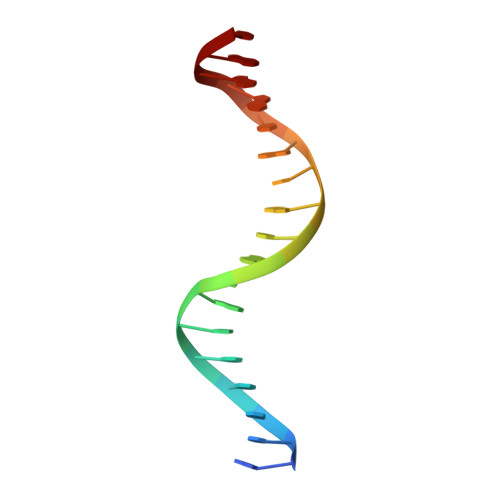
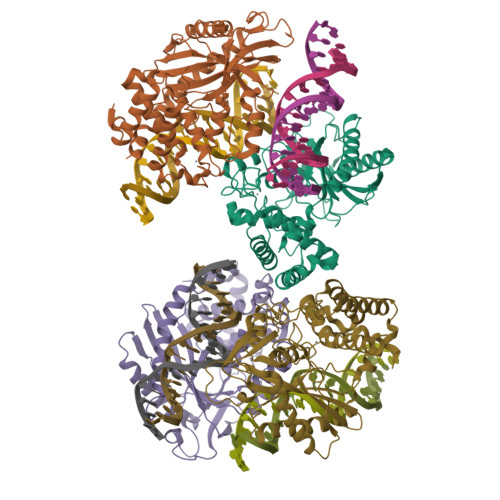
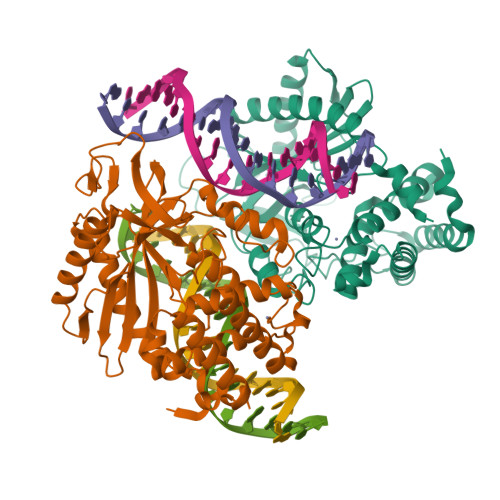
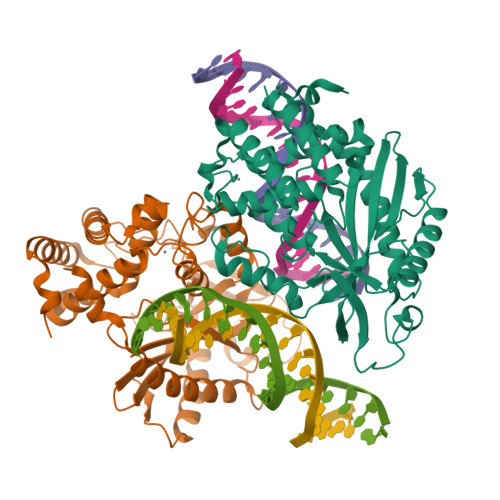
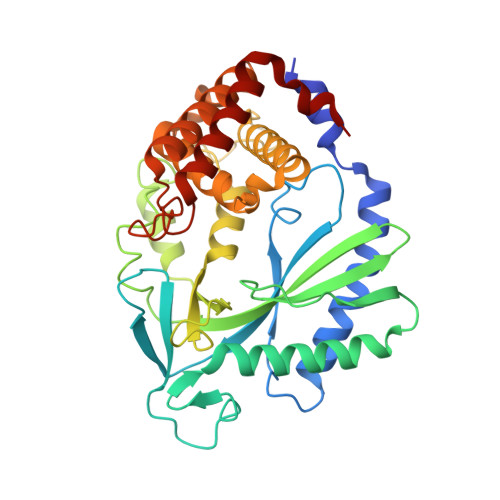
Cyclic GMP-AMP synthase (cGAS) is a cytosolic DNA sensor mediating innate antimicrobial immunity. It catalyzes the synthesis of a noncanonical cyclic dinucleotide, 2',5' cGAMP, that binds to STING and mediates the activation of TBK1 and IRF-3. Activated IRF-3 translocates to the nucleus and initiates the transcription of the IFN-β gene. The structure of mouse cGAS bound to an 18 bp dsDNA revealed that cGAS interacts with dsDNA through two binding sites, forming a 2:2 complex. Enzyme assays and IFN-β reporter assays of cGAS mutants demonstrated that interactions at both DNA binding sites are essential for cGAS activation. Mutagenesis and DNA binding studies showed that the two sites bind dsDNA cooperatively and that site B plays a critical role in DNA binding. The structure of mouse cGAS bound to dsDNA and 2',5' cGAMP provided insight into the catalytic mechanism of cGAS. These results demonstrated that cGAS is activated by dsDNA-induced oligomerization.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843-2128, USA.