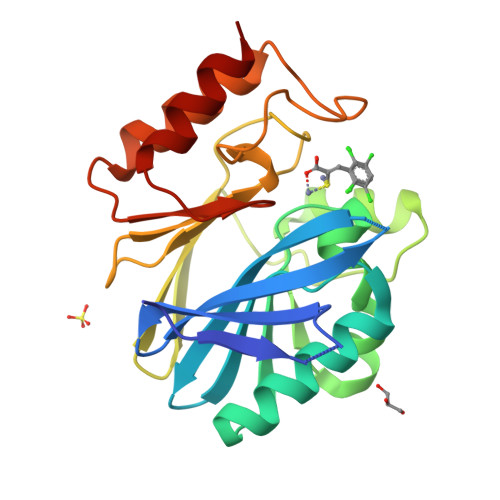
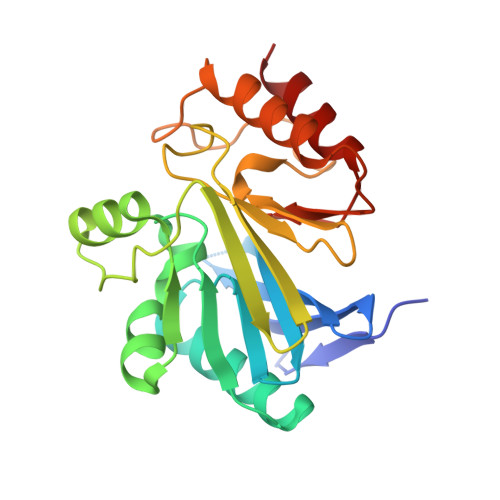
Rhodanine hydrolysis leads to potent thioenolate mediated metallo-beta-lactamase inhibition.
Brem, J., van Berkel, S.S., Aik, W., Rydzik, A.M., Avison, M.B., Pettinati, I., Umland, K.D., Kawamura, A., Spencer, J., Claridge, T.D., McDonough, M.A., Schofield, C.J.(2014) Nat Chem 6: 1084-1090
- PubMed: 25411887
- DOI: https://doi.org/10.1038/nchem.2110
- Primary Citation of Related Structures:
4PVO, 4PVT, 4TYT - PubMed Abstract:
The use of β-lactam antibiotics is compromised by resistance, which is provided by β-lactamases belonging to both metallo (MBL)- and serine (SBL)-β-lactamase subfamilies. The rhodanines are one of very few compound classes that inhibit penicillin-binding proteins (PBPs), SBLs and, as recently reported, MBLs. Here, we describe crystallographic analyses of the mechanism of inhibition of the clinically relevant VIM-2 MBL by a rhodanine, which reveal that the rhodanine ring undergoes hydrolysis to give a thioenolate. The thioenolate is found to bind via di-zinc chelation, mimicking the binding of intermediates in β-lactam hydrolysis. Crystallization of VIM-2 in the presence of the intact rhodanine led to observation of a ternary complex of MBL, a thioenolate fragment and rhodanine. The crystallographic observations are supported by kinetic and biophysical studies, including (19)F NMR analyses, which reveal the rhodanine-derived thioenolate to be a potent broad-spectrum MBL inhibitor and a lead structure for the development of new types of clinically useful MBL inhibitors.
Organizational Affiliation:
Department of Chemistry, University of Oxford, 12 Mansfield Road, Oxford OX1 3TA, UK.