Structural and mechanistic insights into regulation of HBO1 histone acetyltransferase activity by BRPF2.
Tao, Y., Zhong, C., Zhu, J., Xu, S., Ding, J.(2017) Nucleic Acids Res 45: 5707-5719
- PubMed: 28334966
- DOI: https://doi.org/10.1093/nar/gkx142
- Primary Citation of Related Structures:
5GK9 - PubMed Abstract:
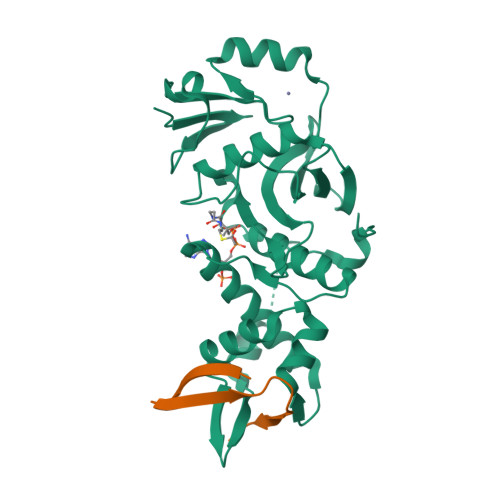
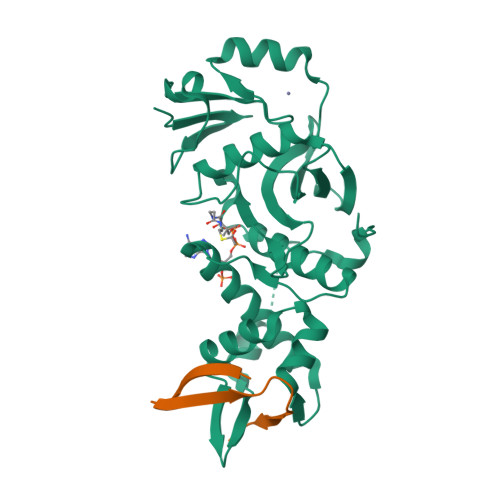
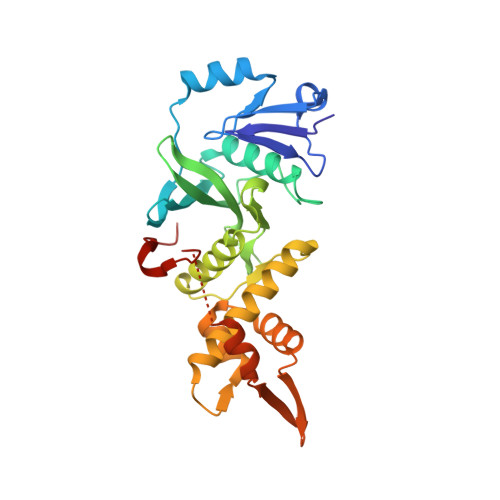
HBO1, a member of the MYST family of histone acetyltransferases (HATs), is required for global acetylation of histone H3K14 and embryonic development. It functions as a catalytic subunit in multisubunit complexes comprising a BRPF1/2/3 or JADE1/2/3 scaffold protein, and two accessory proteins. BRPF2 has been shown to be important for the HAT activity of HBO1 toward H3K14. Here we demonstrated that BRPF2 can regulate the HAT activity of HBO1 toward free H3 and H4, and nucleosomal H3. Particularly, a short N-terminal region of BRPF2 is sufficient for binding to HBO1 and can potentiate its activity toward H3K14. The crystal structure of the HBO1 MYST domain in complex with this segment of BRPF2 together with the biochemical and cell biological data revealed the key residues responsible for the HBO1-BRPF2 interaction. Our structural and functional data together indicate that the N-terminal region of BRPF2 plays an important role in the binding of HBO1 and a minor role in the binding of nucleosomes, which provide new mechanistic insights into the regulation of the HAT activity of HBO1 by BRPF2.
Organizational Affiliation:
National Center for Protein Science Shanghai, State Key Laboratory of Molecular Biology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, and University of Chinese Academy of Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.