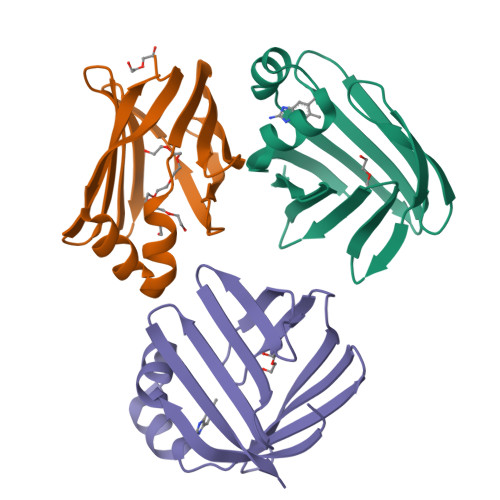
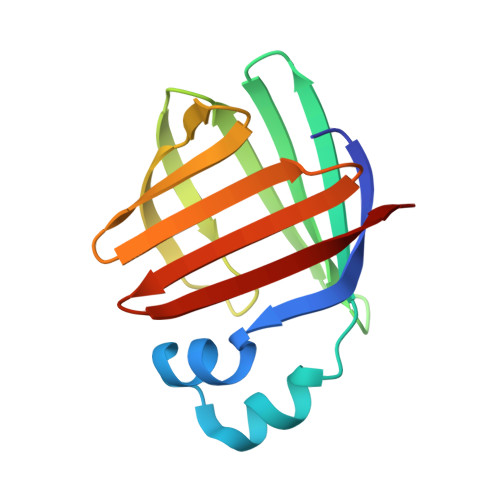
Identification and Investigation of Novel Binding Fragments in the Fatty Acid Binding Protein 6 (FABP6).
Hendrick, A.G., Muller, I., Willems, H., Leonard, P.M., Irving, S., Davenport, R., Ito, T., Reeves, J., Wright, S., Allen, V., Wilkinson, S., Heffron, H., Bazin, R., Turney, J., Mitchell, P.J.(2016) J Med Chem 59: 8094-8102
- PubMed: 27500412
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00869
- Primary Citation of Related Structures:
5L8I, 5L8N, 5L8O - PubMed Abstract:
Fatty acid binding protein 6 (FABP6) is a potential drug discovery target, which, if inhibited, may have a therapeutic benefit for the treatment of diabetes. Currently, there are no published inhibitors of FABP6, and with the target believed to be amenable to fragment-based drug discovery, a structurally enabled program was initiated. This program successfully identified fragment hits using the surface plasmon resonance (SPR) platform. Several hits were validated with SAR and were found to be displaced by the natural ligand taurocholate. We report the first crystal structure of human FABP6 in the unbound form, in complex with cholate, and with one of the key fragments.
Organizational Affiliation:
Takeda Cambridge , 418 Cambridge Science Park, Cambridge CB4 0PZ, United Kingdom.