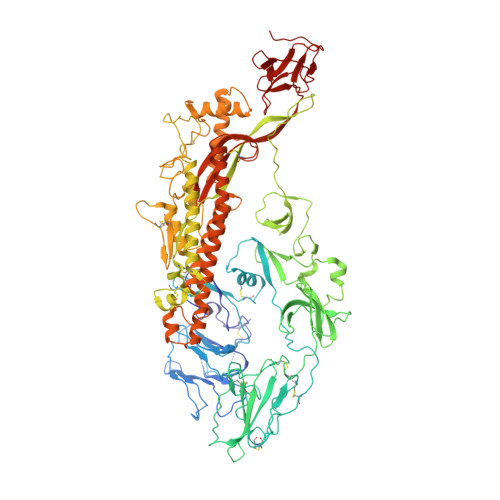
Cryo-electron Microscopy Structure of the Swine Acute Diarrhea Syndrome Coronavirus Spike Glycoprotein Provides Insights into Evolution of Unique Coronavirus Spike Proteins.
Guan, H., Wang, Y., Perculija, V., Saeed, A.F.U.H., Liu, Y., Li, J., Jan, S.S., Li, Y., Zhu, P., Ouyang, S.(2020) J Virol 94
- PubMed: 32817223
- DOI: https://doi.org/10.1128/JVI.01301-20
- Primary Citation of Related Structures:
6M39 - PubMed Abstract:
Coronaviruses (CoV) have caused a number of major epidemics in humans and animals, including the current pandemic of coronavirus disease 2019 (COVID-19), which has brought a renewed focus on the evolution and interspecies transmission of coronaviruses. Swine acute diarrhea syndrome coronavirus (SADS-CoV), which was recently identified in piglets in southern China, is an alphacoronavirus that originates from the same genus of horseshoe bats as severe acute respiratory syndrome CoV (SARS-CoV) and that was reported to be capable of infecting cells from a broad range of species, suggesting a considerable potential for interspecies transmission. Given the importance of the coronavirus spike (S) glycoprotein in host range determination and viral entry, we report a cryo-electron microscopy (cryo-EM) structure of the SADS-CoV S trimer in the prefusion conformation at a 3.55-Å resolution. Our structure reveals that the SADS-CoV S trimer assumes an intrasubunit quaternary packing mode in which the S1 subunit N-terminal domain (S1-NTD) and the S1 subunit C-terminal domain (S1-CTD) of the same protomer pack together by facing each other in the lying-down state. SADS-CoV S has several distinctive structural features that may facilitate immune escape, such as a relatively compact architecture of the S trimer and epitope masking by glycan shielding. Comparison of SADS-CoV S with the spike proteins of the other coronavirus genera suggested that the structural features of SADS-CoV S are evolutionarily related to those of the spike proteins of the other genera rather than to the spike protein of a typical alphacoronavirus. These data provide new insights into the evolutionary relationship between spike glycoproteins of SADS-CoV and those of other coronaviruses and extend our understanding of their structural and functional diversity. IMPORTANCE In this article, we report the atomic-resolution prefusion structure of the spike protein from swine acute diarrhea syndrome coronavirus (SADS-CoV). SADS-CoV is a pathogenic alphacoronavirus that was responsible for a large-scale outbreak of fatal disease in pigs and that was reported to be capable of interspecies transmission. We describe the overall structure of the SADS-CoV spike protein and conducted a detailed analysis of its main structural elements. Our results and analyses are consistent with those of previous phylogenetic studies and suggest that the SADS-CoV spike protein is evolutionarily related to the spike proteins of betacoronaviruses, with a strong similarity in S1-NTDs and a marked divergence in S1-CTDs. Moreover, we discuss the possible immune evasion strategies used by the SADS-CoV spike protein. Our study provides insights into the structure and immune evasion strategies of the SADS-CoV spike protein and broadens the understanding of the evolutionary relationships between coronavirus spike proteins of different genera.
Organizational Affiliation:
Key Laboratory of Innate Immune Biology of Fujian Province, Provincial University Key Laboratory of Cellular Stress Response and Metabolic Regulation, Biomedical Research Center of South China, Key Laboratory of OptoElectronic Science and Technology for Medicine of Ministry of Education, College of Life Sciences, Fujian Normal University, Fuzhou, China.