Nebulized delivery of a broadly neutralizing SARS-CoV-2 RBD-specific nanobody prevents clinical, virological, and pathological disease in a Syrian hamster model of COVID-19.
Esparza, T.J., Chen, Y., Martin, N.P., Bielefeldt-Ohmann, H., Bowen, R.A., Tolbert, W.D., Pazgier, M., Brody, D.L.(null) MAbs 14: 2047144-2047144
- PubMed: 35289719
- DOI: https://doi.org/10.1080/19420862.2022.2047144
- Primary Citation of Related Structures:
7RBY - PubMed Abstract:
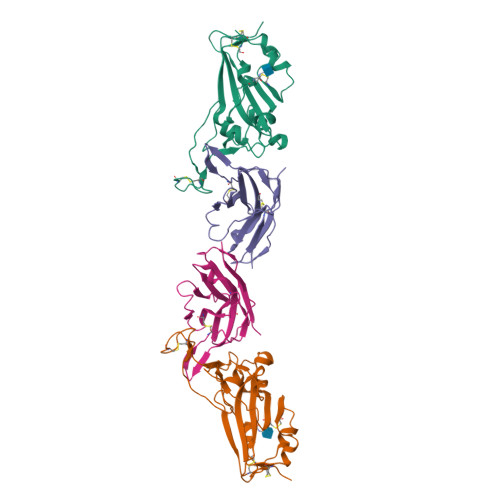
There remains an unmet need for globally deployable, low-cost therapeutics for the ongoing severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic. Previously, we reported on the isolation and in vitro characterization of a potent single-domain nanobody, NIH-CoVnb-112, specific for the receptor-binding domain (RBD) of SARS-CoV-2. Here, we report on the molecular basis for the observed broad in vitro neutralization capability of NIH-CoVnb-112 against variant SARS-CoV-2 pseudoviruses. The structure of NIH-CoVnb-112 bound to SARS-CoV-2 RBD reveals a large contact surface area overlapping the angiotensin converting enzyme 2 (ACE2) binding site, which is largely unencumbered by the common RBD mutations. In an in vivo pilot study, we demonstrate effective reductions in weight loss, viral burden, and lung pathology in a Syrian hamster model of COVID-19 following nebulized delivery of NIH-CoVnb-112. These findings support the further development of NIH-CoVnb-112 as a potential adjunct preventative therapeutic for the treatment of SARS-CoV-2 infection. Abbreviations: ACE2 - angiotensin converting enzyme 2BSA - buried surface areaCDR - complementary determining regionRBD - receptor binding domainRBM - receptor-binding motifSARS-CoV-2 - severe acute respiratory syndrome coronavirus 2.
Organizational Affiliation:
The National Institute of Neurological Disorders and Stroke Intramural Research Program, Laboratory of Functional and Molecular Imaging, Bethesda, MD, USA.