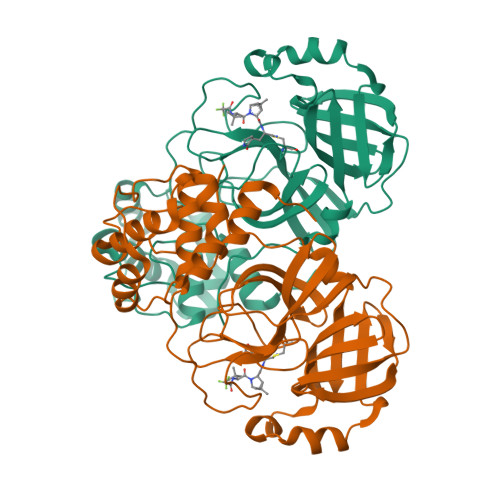
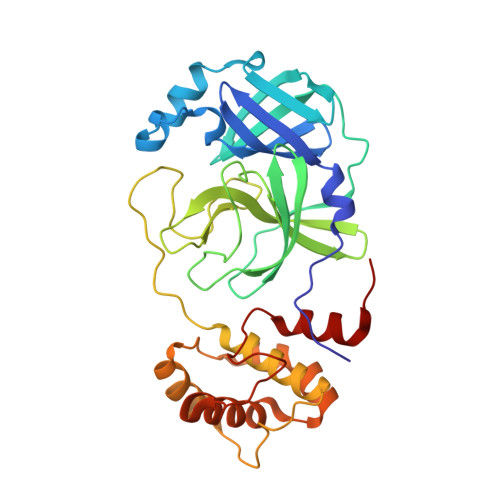
Structural basis for the in vitro efficacy of nirmatrelvir against SARS-CoV-2 variants.
Greasley, S.E., Noell, S., Plotnikova, O., Ferre, R., Liu, W., Bolanos, B., Fennell, K., Nicki, J., Craig, T., Zhu, Y., Stewart, A.E., Steppan, C.M.(2022) J Biological Chem 298: 101972-101972
- PubMed: 35461811
- DOI: https://doi.org/10.1016/j.jbc.2022.101972
- Primary Citation of Related Structures:
7TLL, 7U28, 7U29 - PubMed Abstract:
The COVID-19 pandemic continues to be a public health threat with emerging variants of SARS-CoV-2. Nirmatrelvir (PF-07321332) is a reversible, covalent inhibitor targeting the main protease (M pro ) of SARS-CoV-2 and the active protease inhibitor in PAXLOVID (nirmatrelvir tablets and ritonavir tablets). However, the efficacy of nirmatrelvir is underdetermined against evolving SARS-CoV-2 variants. Here, we evaluated the in vitro catalytic activity and potency of nirmatrelvir against the M pro of prevalent variants of concern (VOCs) or variants of interest (VOIs): Alpha (α, B.1.1.7), Beta (β, B.1.351), Delta (δ, B1.617.2), Gamma (γ, P.1), Lambda (λ, B.1.1.1.37/C37), Omicron (ο, B.1.1.529), as well as the original Washington or wildtype strain. These VOCs/VOIs carry prevalent mutations at varying frequencies in the M pro specifically for α, β, γ (K90R), λ (G15S), and ο (P132H). In vitro biochemical enzymatic assay characterization of the enzyme kinetics of the mutant M pros demonstrates that they are catalytically comparable to wildtype. We found that nirmatrelvir has similar potency against each mutant M pro including P132H that is observed in the Omicron variant with a Ki of 0.635 nM as compared to a Ki of 0.933 nM for wildtype. The molecular basis for these observations were provided by solution-phase structural dynamics and structural determination of nirmatrelvir bound to the ο, λ, and β M pro at 1.63 to 2.09 Å resolution. These in vitro data suggest that PAXLOVID has the potential to maintain plasma concentrations of nirmatrelvir many-fold times higher than the amount required to stop the SARS-CoV-2 VOC/VOI, including Omicron, from replicating in cells.
Organizational Affiliation:
Medicine Design, Pfizer Worldwide Research, Development & Medical, La Jolla, California, USA.