Broadly potent anti-SARS-CoV-2 antibody shares 93% of epitope with ACE2 and provides full protection in monkeys.
Fenwick, C., Turelli, P., Duhoo, Y., Lau, K., Herate, C., Marlin, R., Lamrayah, M., Campos, J., Esteves-Leuenberger, L., Farina, A., Raclot, C., Genet, V., Fiscalini, F., Cesborn, J., Perez, L., Dereuddre-Bosquet, N., Contreras, V., Lheureux, K., Relouzat, F., Abdelnabi, R., Leyssen, P., Levy, Y., Pojer, F., Le Grand, R., Trono, D., Pantaleo, G.(2023) J Infect 87: 524-537
- PubMed: 37852477
- DOI: https://doi.org/10.1016/j.jinf.2023.10.008
- Primary Citation of Related Structures:
8PQ2, 8PSD - PubMed Abstract:
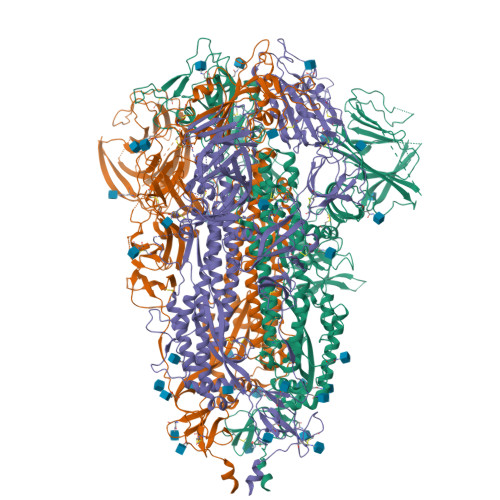
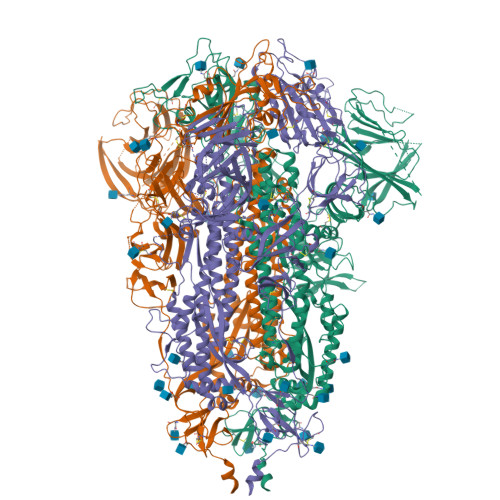
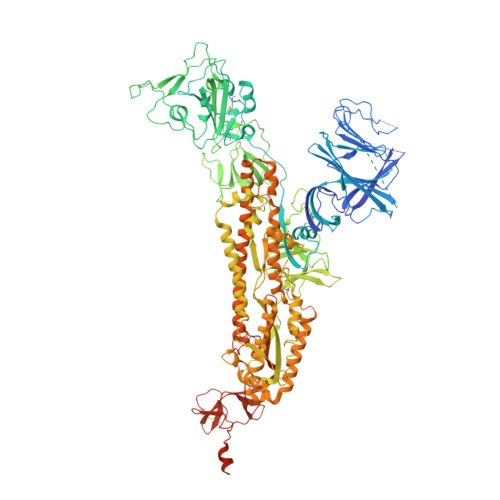
Due to the rapid evolution of SARS-CoV-2 to variants with reduced sensitivity to vaccine-induced humoral immunity and the near complete loss of protective efficacy of licensed therapeutic monoclonal antibodies, we isolated a potent, broad-spectrum neutralizing antibody that could potentially provide prophylactic protection to immunocompromised patient populations. Spike-specific B-cell clones isolated from a vaccinated post-infected donor were profiled for those producing potent neutralizing antibodies against a panel of SARS-CoV-2 variants. The P4J15 antibody was further characterized to define the structural binding epitope, viral resistance, and in vivo efficacy. The P4J15 mAb shows <20 ng/ml neutralizing activity against all variants including the latest XBB.2.3 and EG.5.1 sub-lineages. Structural studies of P4J15 in complex with Omicron XBB.1 Spike show that the P4J15 epitope shares ∼93% of its buried surface area with the ACE2 contact region, consistent with an ACE2 mimetic antibody. In vitro selection of SARS-CoV-2 mutants escaping P4J15 neutralization showed reduced infectivity, poor ACE2 binding, and mutations are rare in public sequence databases. Using a SARS-CoV-2 XBB.1.5 monkey challenge model, P4J15-LS confers complete prophylactic protection with an exceptionally long in vivo half-life of 43 days. The P4J15 mAb has potential as a broad-spectrum anti-SARS-CoV-2 drug for prophylactic protection of at-risk patient populations.
Organizational Affiliation:
Service of Immunology and Allergy, Department of Medicine, Lausanne University Hospital and University of Lausanne, Lausanne, Switzerland.