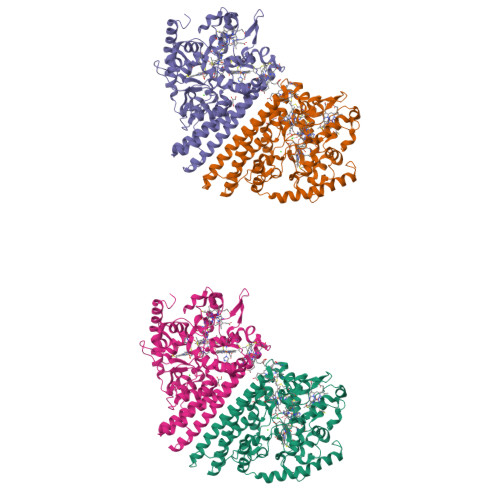
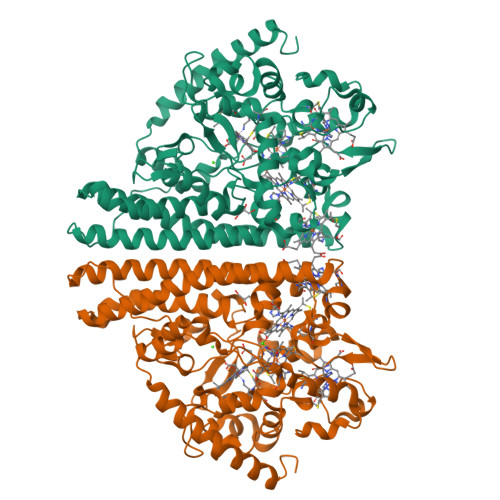
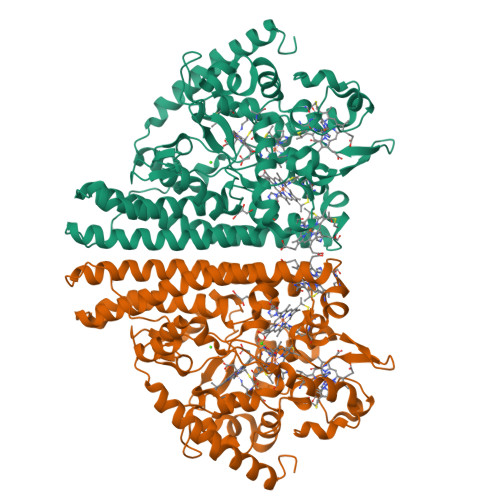
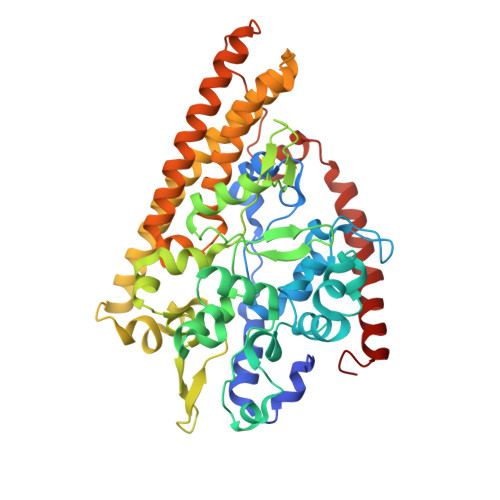
Structure and Spectroscopy of the Periplasmic Cytochrome C Nitrite Reductase from Escherichia Coli
Bamford, V.A., Angove, H.C., Seward, H.E., Thomson, A.J., Cole, J.A., Butt, J.N., Hemmings, A.M., Richardson, D.J.(2002) Biochemistry 41: 2921
- PubMed: 11863430
- DOI: https://doi.org/10.1021/bi015765d
- Primary Citation of Related Structures:
1GU6 - PubMed Abstract:
The crystal structure and spectroscopic properties of the periplasmic penta-heme cytochrome c nitrite reductase (NrfA) of Escherichia coli are presented. The structure is the first for a member of the NrfA subgroup that utilize a soluble penta-heme cytochrome, NrfB, as a redox partner. Comparison to the structures of Wolinella succinogenes NrfA and Sulfospirillum deleyianum NrfA, which accept electrons from a membrane-anchored tetra-heme cytochrome (NrfH), reveals notable differences in the protein surface around heme 2, which may be the docking site for the redox partner. The structure shows that four of the NrfA hemes (hemes 2-5) have bis-histidine axial heme-Fe ligation. The catalytic heme-Fe (heme 1) has a lysine distal ligand and an oxygen atom proximal ligand. Analysis of NrfA in solution by magnetic circular dichroism (MCD) suggested that the oxygen ligand arose from water. Electron paramagnetic resonance (EPR) spectra were collected from electrochemically poised NrfA samples. Broad perpendicular mode signals at g similar 10.8 and 3.5, characteristic of weakly spin-coupled S = 5/2, S = 1/2 paramagnets, titrated with E(m) = -107 mV. A possible origin for these are the active site Lys-OH(2) coordinated heme (heme 1) and a nearby bis-His coordinated heme (heme 3). A rhombic heme Fe(III) EPR signal at g(z) = 2.91, g(y) = 2.3, g(x) = 1.5 titrated with E(m) = -37 mV and is likely to arise from bis-His coordinated heme (heme 2) in which the interplanar angle of the imidazole rings is 21.2. The final two bis-His coordinated hemes (hemes 4 and 5) have imidazole interplanar angles of 64.4 and 71.8. Either, or both, of these hemes could give rise to a "Large g max" EPR signal at g(z)() = 3.17 that titrated at potentials between -250 and -400 mV. Previous spectroscopic studies on NrfA from a number of bacterial species are considered in the light of the structure-based spectro-potentiometric analysis presented for the E. coli NrfA.
Organizational Affiliation:
Centre for Metalloprotein Spectroscopy and Biology, School of Biological Sciences, University of East Anglia, Norwich, NR4 7TJ, United Kingdom.