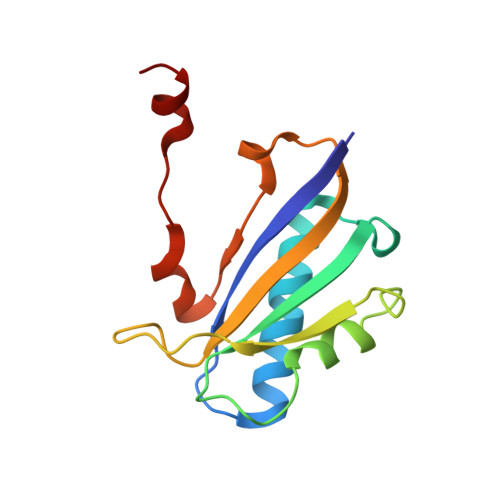
The 1.30 A resolution structure of the Bacillus subtilis chorismate mutase catalytic homotrimer.
Ladner, J.E., Reddy, P., Davis, A., Tordova, M., Howard, A.J., Gilliland, G.L.(2000) Acta Crystallogr D Biol Crystallogr 56: 673-683
- PubMed: 10818343
- DOI: https://doi.org/10.1107/s0907444900004625
- Primary Citation of Related Structures:
1DBF - PubMed Abstract:
The crystal structure of the Bacillus subtilis chorismate mutase, an enzyme of the aromatic amino acids biosynthetic pathway, was determined to 1.30 A resolution. The structure of the homotrimer was determined by molecular replacement using orthorhombic crystals of space group P2(1)2(1)2(1) with unit-cell parameters a = 52.2, b = 83. 8, c = 86.0 A. The ABC trimer of the monoclinic crystal structure [Chook et al. (1994), J. Mol. Biol. 240, 476-500] was used as the starting model. The final coordinates are composed of three complete polypeptide chains of 127 amino-acid residues. In addition, there are nine sulfate ions, five glycerol molecules and 424 water molecules clearly visible in the structure. This structure was refined with aniosotropic temperature factors, has excellent geometry and a crystallographic R factor of 0.169 with an R(free) of 0.236. The three active sites of the macromolecule are at the subunit interfaces, with residues from two subunits contributing to each site. This orthorhombic crystal form was grown using ammonium sulfate as the precipitant; glycerol was used as a cryoprotectant during data collection. A glycerol molecule and sulfate ion in each of the active sites was found mimicking a transition-state analog. In this structure, the C-terminal tails of the subunits of the trimer are hydrogen bonded to residues of the active site of neighboring trimers in the crystal and thus cross-link the molecules in the crystal lattice.
Organizational Affiliation:
National Institute of Standards and Technology and the University of Maryland Biotechnology Institute, Rockville, MD 20850, USA.