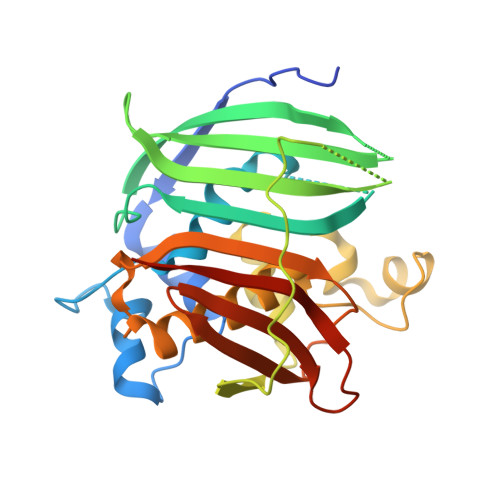
Crystal structure of 2-enoyl-CoA hydratase 2 from human peroxisomal multifunctional enzyme type 2.
Koski, M.K., Haapalainen, A.M., Hiltunen, J.K.(2005) J Mol Biol 345: 1157-1169
- PubMed: 15644212
- DOI: https://doi.org/10.1016/j.jmb.2004.11.009
- Primary Citation of Related Structures:
1S9C - PubMed Abstract:
2-Enoyl-CoA hydratase 2 is the middle part of the mammalian peroxisomal multifunctional enzyme type 2 (MFE-2), which is known to be important in the beta-oxidation of very-long-chain and alpha-methyl-branched fatty acids as well as in the synthesis of bile acids. Here, we present the crystal structure of the hydratase 2 from the human MFE-2 to 3A resolution. The three-dimensional structure resembles the recently solved crystal structure of hydratase 2 from the yeast, Candida tropicalis, MFE-2 having a two-domain subunit structure with a C-domain complete hot-dog fold housing the active site, and an N-domain incomplete hot-dog fold housing the cavity for the aliphatic acyl part of the substrate molecule. The ability of human hydratase 2 to utilize such bulky compounds which are not physiological substrates for the fungal ortholog, e.g. CoA esters of C26 fatty acids, pristanic acid and di/trihydroxycholestanoic acids, is explained by a large hydrophobic cavity formed upon the movements of the extremely mobile loops I-III in the N-domain. In the unliganded form of human hydratase 2, however, the loop I blocks the entrance of fatty enoyl-CoAs with chain-length >C8. Therefore, we expect that upon binding of substrates bulkier than C8, the loop I gives way, contemporaneously causing a secondary effect in the CoA-binding pocket and/or active site required for efficient hydration reaction. This structural feature would explain the inactivity of human hydratase 2 towards short-chain substrates. The solved structure is also used as a tool for analyzing the various inactivating mutations, identified among others in MFE-2-deficient patients. Since hydratase 2 is the last functional unit of mammalian MFE-2 whose structure has been solved, the organization of the functional units in the biologically active full-length enzyme is also discussed.
Organizational Affiliation:
Department of Biochemistry and Biocenter Oulu, University of Oulu, Box 3000, FIN-90014 Oulu, Finland.