A Simple Protocol To Estimate Differences in Protein Binding Affinity for Enantiomers without Prior Resolution of Racemates
Fokkens, J., Klebe, G.(2006) Angew Chem Int Ed Engl 45: 985-989
Experimental Data Snapshot
(2006) Angew Chem Int Ed Engl 45: 985-989
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Prothrombin light chain | A [auth L] | 27 | Homo sapiens | Mutation(s): 0 EC: 3.4.21.5 | 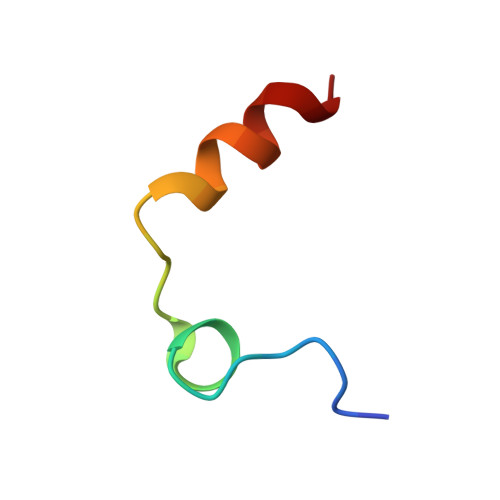 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00734 (Homo sapiens) Explore P00734 Go to UniProtKB: P00734 | |||||
PHAROS: P00734 GTEx: ENSG00000180210 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00734 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Prothrombin heavy chain | B [auth H] | 257 | Homo sapiens | Mutation(s): 0 EC: 3.4.21.5 | 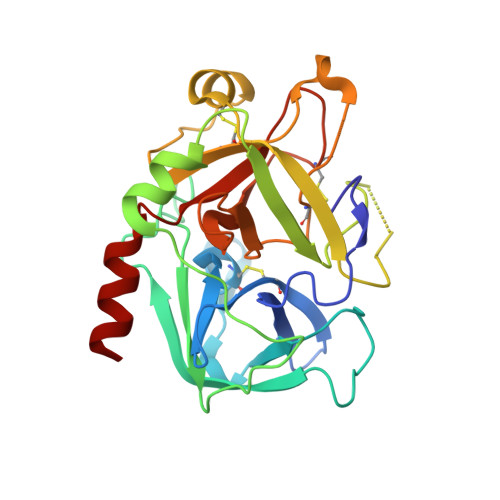 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00734 (Homo sapiens) Explore P00734 Go to UniProtKB: P00734 | |||||
PHAROS: P00734 GTEx: ENSG00000180210 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00734 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 10-mer peptide from Acety-Hirudin(54-65) sulfated | C [auth I] | 10 | Hirudo medicinalis | Mutation(s): 0 | 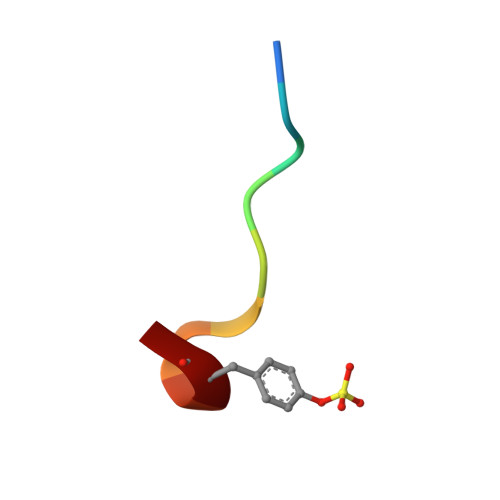 |
UniProt | |||||
Find proteins for P01050 (Hirudo medicinalis) Explore P01050 Go to UniProtKB: P01050 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01050 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CCR Query on CCR | D [auth H] | [N-[N-(4-METHOXY-2,3,6-TRIMETHYLPHENYLSULFONYL)-L-ASPARTYL]-D-(4-AMIDINO-PHENYLALANYL)]-PIPERIDINE C29 H39 N5 O7 S ZOXOKTJHZSUHRJ-XZOQPEGZSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| TYS Query on TYS | C [auth I] | L-PEPTIDE LINKING | C9 H11 N O6 S |  | TYR |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000628 (CCR) Query on PRD_000628 | D [auth H] | [N-[N-(4-METHOXY-2,3,6-TRIMETHYLPHENYLSULFONYL)-L-ASPARTYL]-D-(4-AMIDINO-PHENYLALANYL)]-PIPERIDINE | Peptide-like / Trypsin inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 70.885 | α = 90 |
| b = 71.744 | β = 100.93 |
| c = 72.274 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SHELX | model building |
| SHELXL-97 | refinement |
| SCALEPACK | data scaling |
| CNS | phasing |