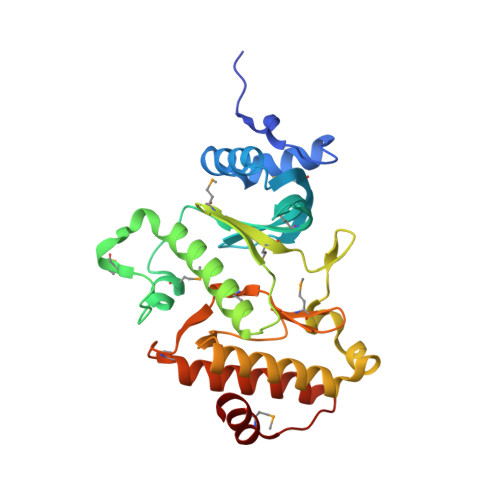
Structure and activity of the atypical serine kinase Rio1.
Laronde-Leblanc, N., Guszczynski, T., Copeland, T., Wlodawer, A.(2005) FEBS J 272: 3698-3713
- PubMed: 16008568
- DOI: https://doi.org/10.1111/j.1742-4658.2005.04796.x
- Primary Citation of Related Structures:
1ZP9, 1ZTF, 1ZTH - PubMed Abstract:
Rio1 is the founding member of the RIO family of atypical serine kinases that are universally present in all organisms from archaea to mammals. Activity of Rio1 was shown to be absolutely essential in Saccharomyces cerevisiae for the processing of 18S ribosomal RNA, as well as for proper cell cycle progression and chromosome maintenance. We determined high-resolution crystal structures of Archaeoglobus fulgidus Rio1 in the presence and absence of bound nucleotides. Crystallization of Rio1 in the presence of ATP or ADP and manganese ions demonstrated major conformational changes in the active site, compared with the uncomplexed protein. Comparisons of the structure of Rio1 with the previously determined structure of the Rio2 kinase defined the minimal RIO domain and the distinct features of the RIO subfamilies. We report here that Ser108 represents the sole autophosphorylation site of A. fulgidus Rio1 and have therefore established its putative peptide substrate. In addition, we show that a mutant enzyme that cannot be autophosphorylated can still phosphorylate an inactive form of Rio1, as well as a number of typical kinase substrates.
Organizational Affiliation:
Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute, NCI-Frederick, MD 21702-1201, USA.