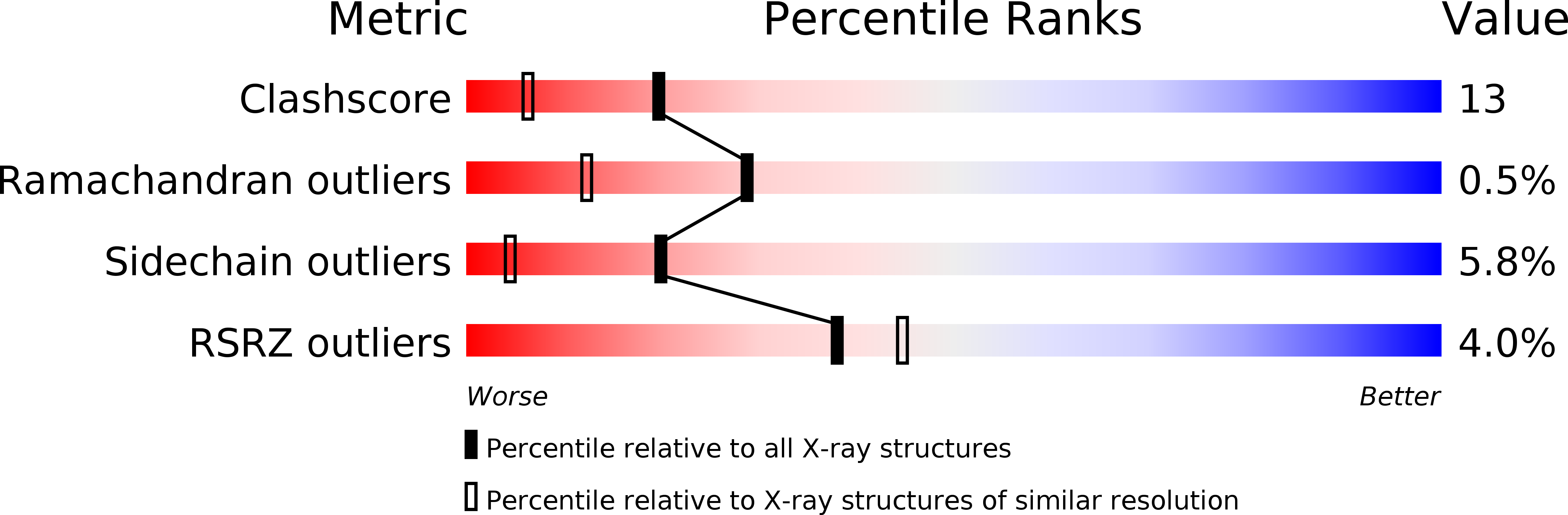
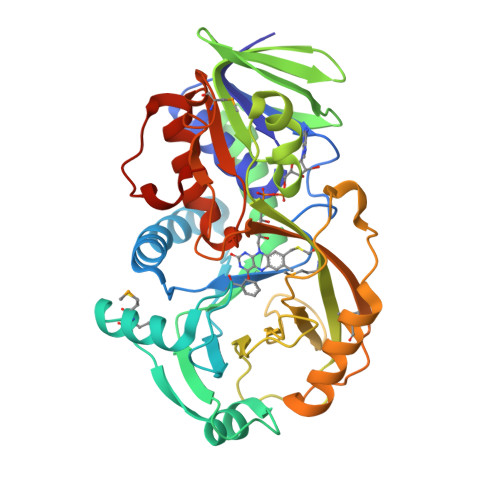
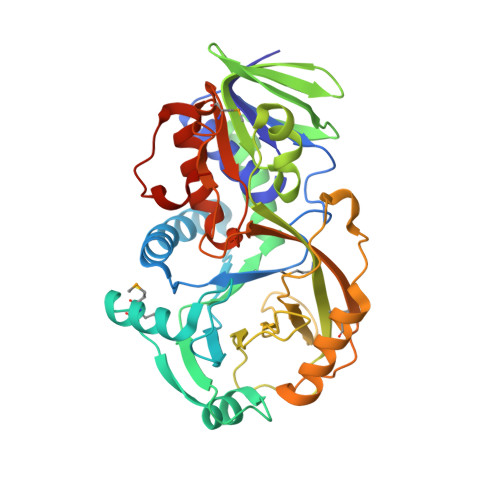
NikD, an Unusual Amino Acid Oxidase Essential for Nikkomycin Biosynthesis: Structures of Closed and Open Forms at 1.15 and 1.90 A Resolution
Carrell, C.J., Bruckner, R.C., Venci, D., Zhao, G., Jorns, M.S., Mathews, F.S.(2007) Structure 15: 928-941
- PubMed: 17697998
- DOI: https://doi.org/10.1016/j.str.2007.06.010
- Primary Citation of Related Structures:
2OLN, 2OLO, 2Q6U - PubMed Abstract:
NikD is an unusual amino-acid-oxidizing enzyme that contains covalently bound FAD, catalyzes a 4-electron oxidation of piperideine-2-carboxylic acid to picolinate, and plays a critical role in the biosynthesis of nikkomycin antibiotics. Crystal structures of closed and open forms of nikD, a two-domain enzyme, have been determined to resolutions of 1.15 and 1.9 A, respectively. The two forms differ by an 11 degrees rotation of the catalytic domain with respect to the FAD-binding domain. The active site is inaccessible to solvent in the closed form; an endogenous ligand, believed to be picolinate, is bound close to and parallel with the flavin ring, an orientation compatible with redox catalysis. The active site is solvent accessible in the open form, but the picolinate ligand is approximately perpendicular to the flavin ring and a tryptophan is stacked above the flavin ring. NikD also contains a mobile cation binding loop.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110, USA.