Molecular Mechanisms of Yeast Cell Wall Glucan Remodeling.
Hurtado-Guerrero, R., Schuttelkopf, A.W., Mouyna, I., Ibrahim, A.F.M., Shepherd, S., Fontaine, T., Latge, J., Van Aalten, D.M.F.(2009) J Biological Chem 284: 8461
- PubMed: 19097997
- DOI: https://doi.org/10.1074/jbc.M807990200
- Primary Citation of Related Structures:
2W61, 2W62, 2W63 - PubMed Abstract:
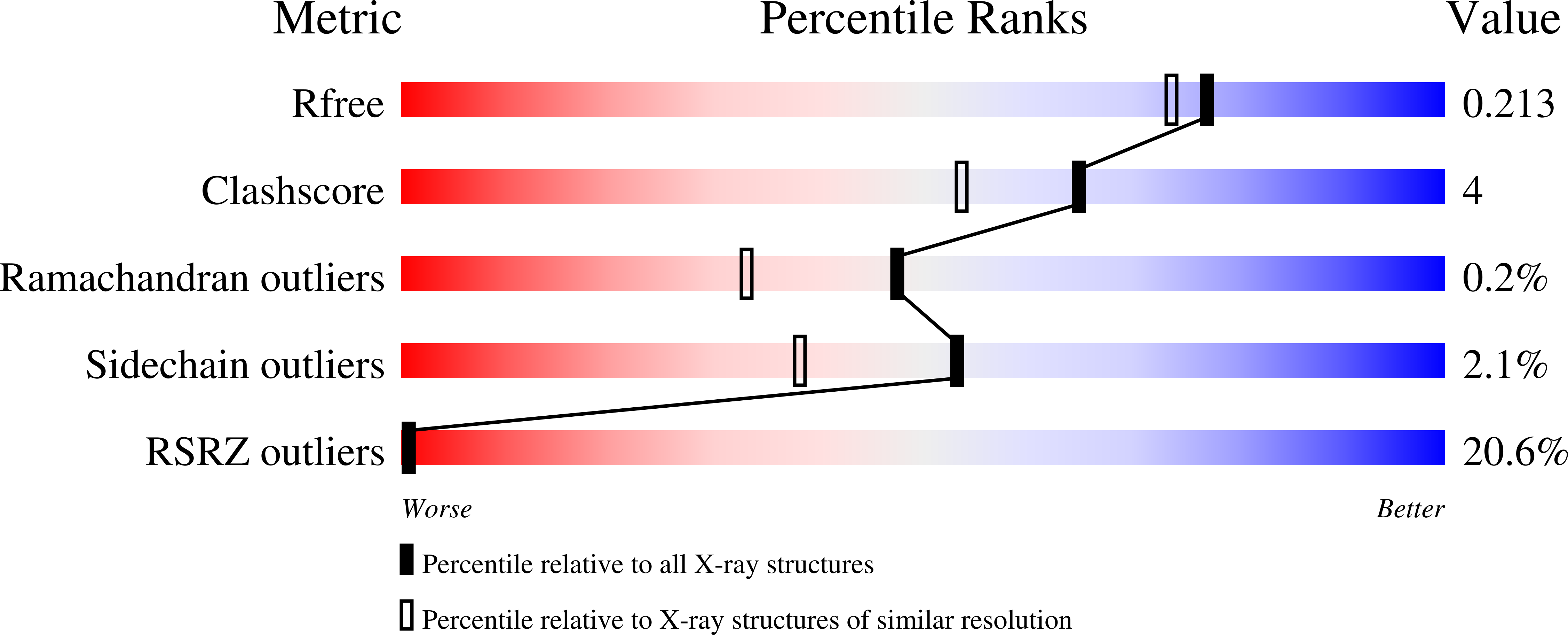
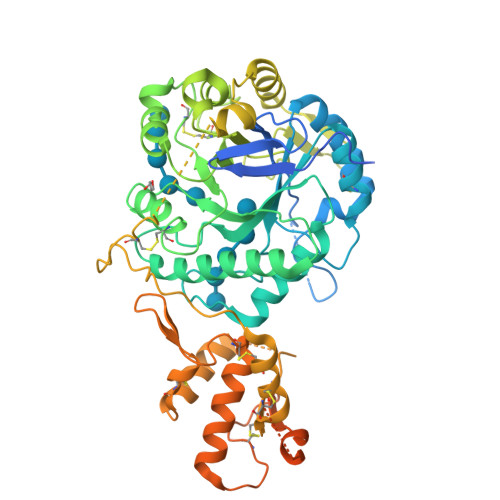
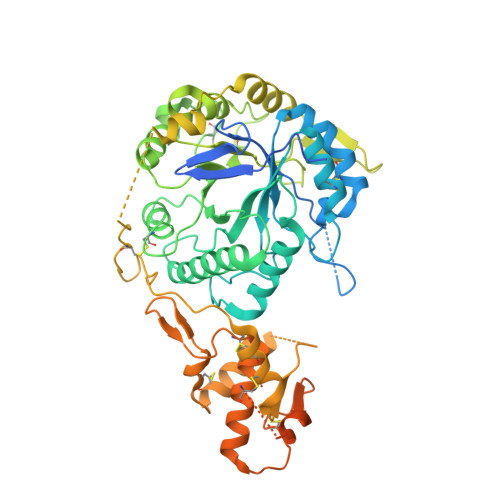
Yeast cell wall remodeling is controlled by the equilibrium between glycoside hydrolases, glycosyltransferases, and transglycosylases. Family 72 glycoside hydrolases (GH72) are ubiquitous in fungal organisms and are known to possess significant transglycosylase activity, producing elongated beta(1-3) glucan chains. However, the molecular mechanisms that control the balance between hydrolysis and transglycosylation in these enzymes are not understood. Here we present the first crystal structure of a glucan transglycosylase, Saccharomyces cerevisiae Gas2 (ScGas2), revealing a multidomain fold, with a (betaalpha)(8) catalytic core and a separate glucan binding domain with an elongated, conserved glucan binding groove. Structures of ScGas2 complexes with different beta-glucan substrate/product oligosaccharides provide "snapshots" of substrate binding and hydrolysis/transglycosylation giving the first insights into the mechanisms these enzymes employ to drive beta(1-3) glucan elongation. Together with mutagenesis and analysis of reaction products, the structures suggest a "base occlusion" mechanism through which these enzymes protect the covalent protein-enzyme intermediate from a water nucleophile, thus controlling the balance between hydrolysis and transglycosylation and driving the elongation of beta(1-3) glucan chains in the yeast cell wall.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, United Kingdom. R.Hurtadoguerrero@dundee.ac.uk