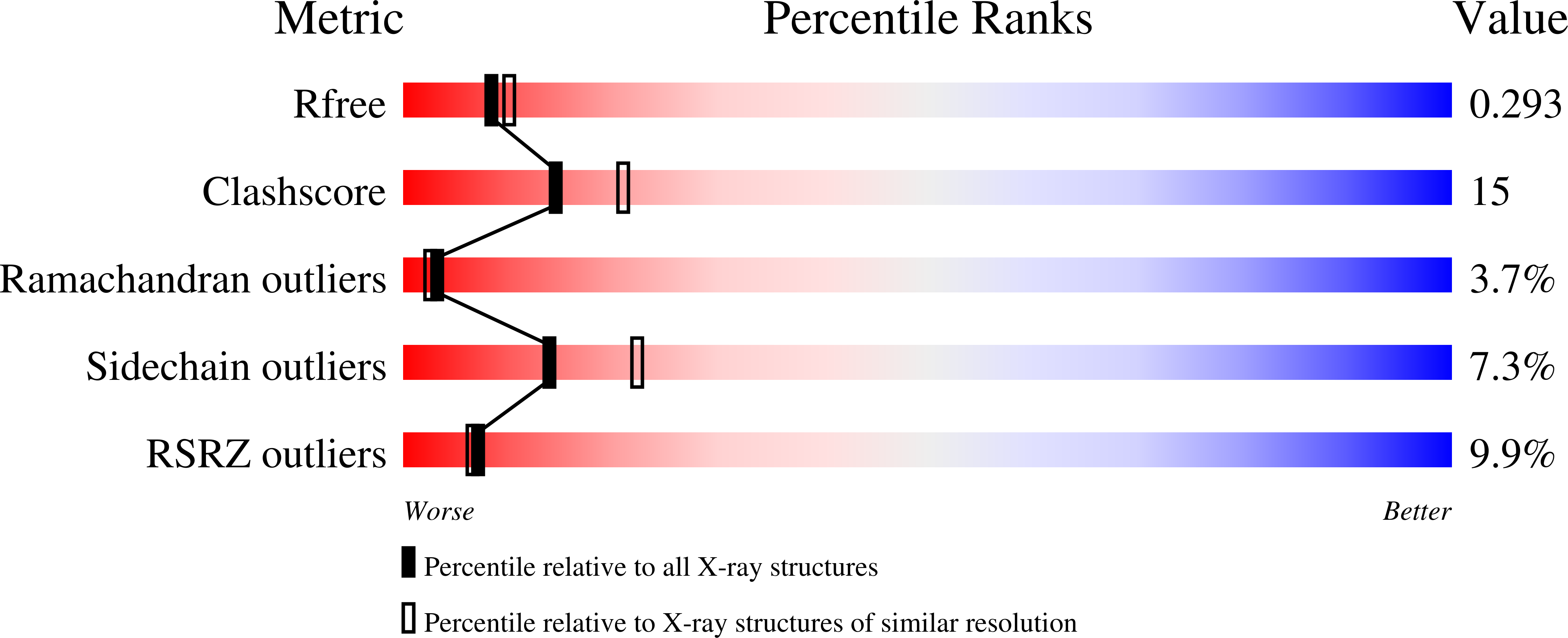
Structural and Mutational Analyses of a CD8alphabeta Heterodimer and Comparison with the CD8alphaalpha Homodimer.
Chang, H.C., Tan, K., Ouyang, J., Parisini, E., Liu, J.H., Le, Y., Wang, X., Reinherz, E.L., Wang, J.H.(2005) Immunity 23: 661-671
- PubMed: 16356863
- DOI: https://doi.org/10.1016/j.immuni.2005.11.002
- Primary Citation of Related Structures:
2ATP - PubMed Abstract:
The crystal structure of a recombinant mouse single chain CD8alphabeta ectodomains at 2.4 A resolution reveals paired immunoglobulin variable region-like domains with a striking resemblance to CD8alphaalpha in size, shape, and surface electrostatic potential of complementarity-determining regions (CDR), despite <20% sequence identity between the CD8alpha and CD8beta subunits. Unlike the CD8alpha subunit(s) in the heterodimer or homodimer, the CDR1 loop of CD8beta tilts away from its corresponding CDR2 and CDR3 loops. Consistent with this observation, independent mutational studies reveal that alanine substitutions of residues in the CDR1 loop of CD8beta have no effect on CD8alphabeta coreceptor function, whereas mutations in CD8beta CDR2 and CDR3 loops abolish CD8alphabeta coreceptor activity. The implications of these findings and additional CD8alpha mutational studies for CD8alphabeta- versus CD8alphaalpha-MHCI binding are discussed.
Organizational Affiliation:
Laboratory of Immunobiology, Department of Medical Oncology, Dana-Farber Cancer Institute, Department of Medicine, Harvard Medical School, Boston, Massachusetts 02115, USA. hsiu-ching_chang@dfci.harvard.edu