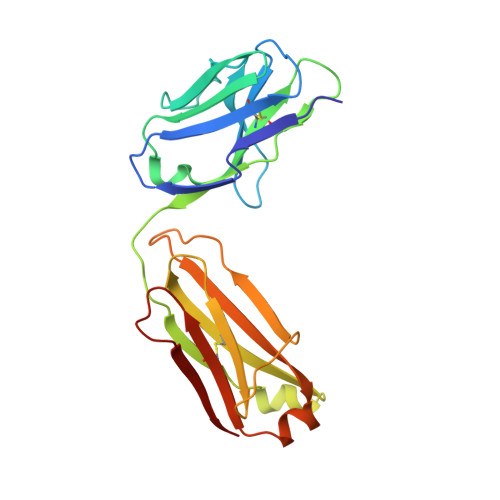
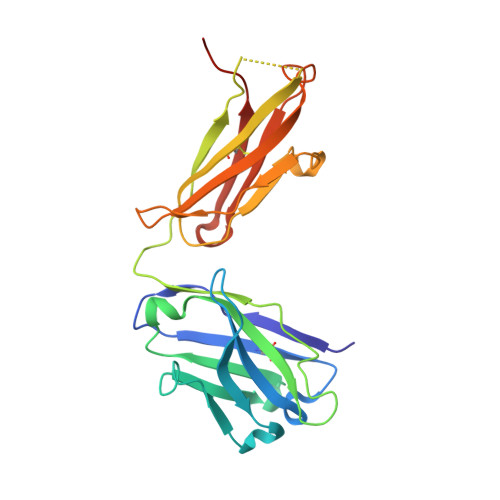
Structure-function studies of two synthetic anti-vascular endothelial growth factor Fabs and comparison with the Avastin Fab.
Fuh, G., Wu, P., Liang, W.C., Ultsch, M., Lee, C.V., Moffat, B., Wiesmann, C.(2006) J Biol Chem 281: 6625-6631
- PubMed: 16373345
- DOI: https://doi.org/10.1074/jbc.M507783200
- Primary Citation of Related Structures:
2FJF, 2FJG, 2FJH - PubMed Abstract:
In the quest to discover new research tools and to develop better agents in the fight against cancer, two antibodies, G6 and B20-4, were isolated from synthetic antibody phage libraries. Unlike the AVASTINtrade mark antibody, a recently approved agent for the treatment of patients with colorectal cancer, B20-4 and G6 bind and block both human and murine vascular endothelial growth factor (VEGF). Here we have analyzed and compared the binding epitopes on VEGF for these three antibodies using alanine-scanning mutagenesis and structural analyses. The epitopes recognized by both synthetic antibodies are conserved between human and mouse VEGF, and they match closely to the receptor epitopes both structurally and functionally. In contrast, the Avastin epitope overlaps minimally with the receptor binding surface and centers around a residue that is not conserved in mouse. Our structural and functional analyses elucidate the cross-species reactivity of all three antibodies and emphasize the potential advantages of antibody generation using phage display as the resulting antibodies do not depend on sequence differences across species and preferentially target natural protein-protein interaction surfaces.
Organizational Affiliation:
Department of Protein Engineering, Genentech, Inc., South San Francisco, California 94080, USA. gml@gene.com