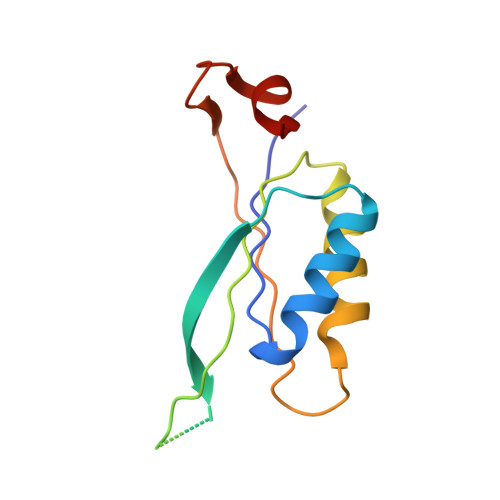
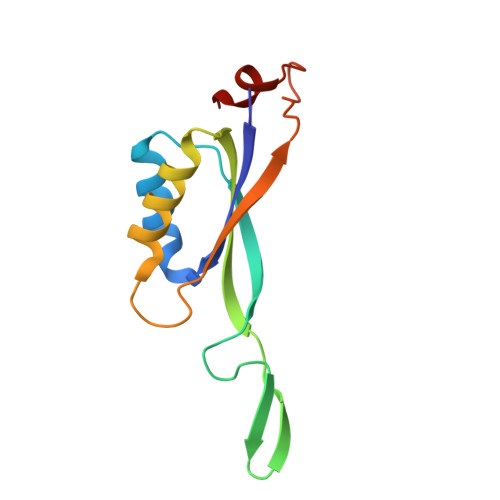
Structure of Glnk1 with Bound Effectors Indicates Regulatory Mechanism for Ammonia Uptake.
Yildiz, O., Kalthoff, C., Raunser, S., Kuhlbrandt, W.(2007) EMBO J 26: 589
- PubMed: 17203075
- DOI: https://doi.org/10.1038/sj.emboj.7601492
- Primary Citation of Related Structures:
2J9C, 2J9D, 2J9E - PubMed Abstract:
A binary complex of the ammonia channel Amt1 from Methanococcus jannaschii and its cognate P(II) signalling protein GlnK1 has been produced and characterized. Complex formation is prevented specifically by the effector molecules Mg-ATP and 2-ketoglutarate. Single-particle electron microscopy of the complex shows that GlnK1 binds on the cytoplasmic side of Amt1. Three high-resolution X-ray structures of GlnK1 indicate that the functionally important T-loop has an extended, flexible conformation in the absence of Mg-ATP, but assumes a compact, tightly folded conformation upon Mg-ATP binding, which in turn creates a 2-ketoglutarate-binding site. We propose a regulatory mechanism by which nitrogen uptake is controlled by the binding of both effector molecules to GlnK1. At normal effector levels, a 2-ketoglutarate molecule binding at the apex of the compact T-loop would prevent complex formation, ensuring uninhibited ammonia uptake. At low levels of Mg-ATP, the extended loops would seal the ammonia channels in the complex. Binding of both effector molecules to P(II) signalling proteins may thus represent an effective feedback mechanism for regulating ammonium uptake through the membrane.
Organizational Affiliation:
Department of Structural Biology, Max Planck Institute of Biophysics, Frankfurt am Main, Germany.