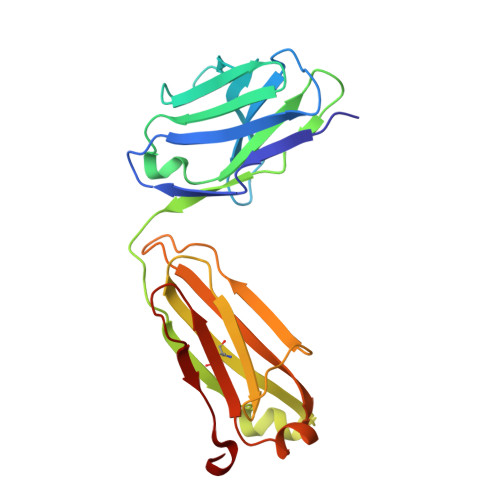
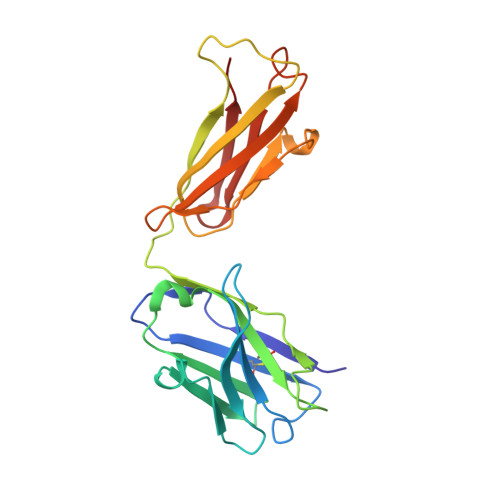
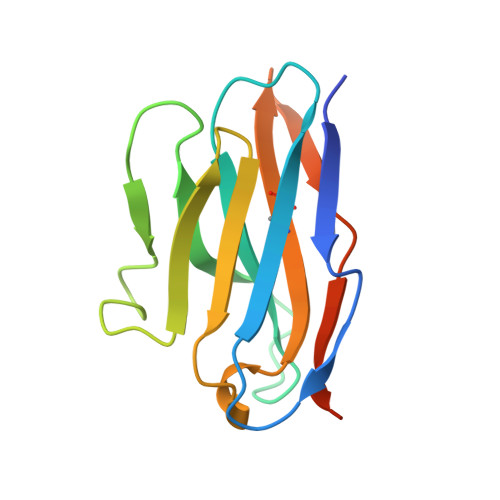
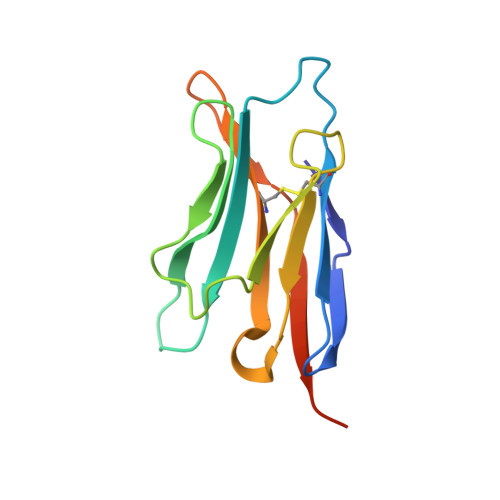
The Crystal Structure of CD8 in Complex with YTS156.7.7 Fab and Interaction with Other CD8 Antibodies Define the Binding Mode of CD8 alphabeta to MHC Class I
Shore, D.A., Issafras, H., Landais, E., Teyton, L., Wilson, I.A.(2008) J Mol Biology 384: 1190-1202
- PubMed: 18929574
- DOI: https://doi.org/10.1016/j.jmb.2008.09.069
- Primary Citation of Related Structures:
3B9K - PubMed Abstract:
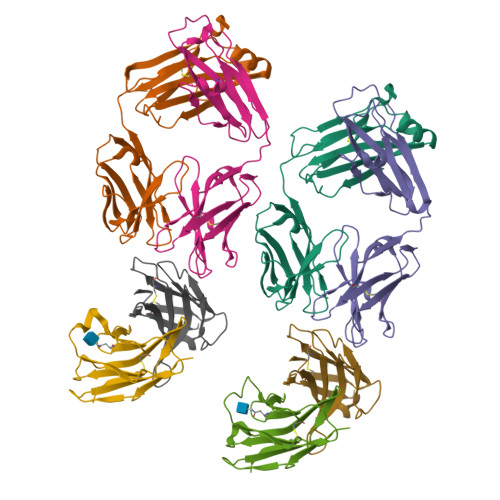
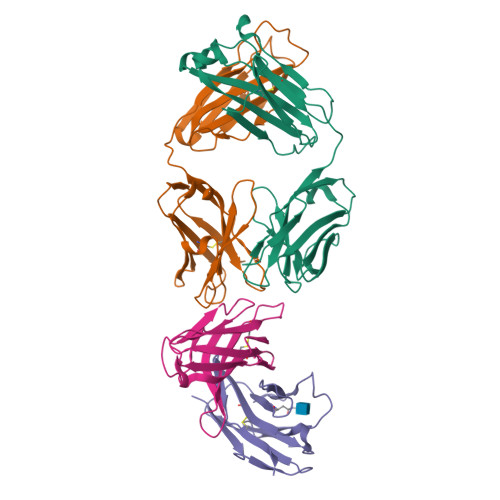
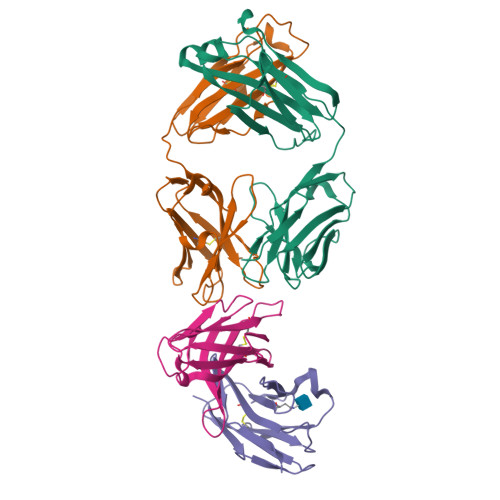
The CD8alphabeta heterodimer interacts with class I pMHC on antigen-presenting cells as a co-receptor for TCR-mediated activation of cytotoxic T cells. To characterize this immunologically important interaction, we used monoclonal antibodies (mAbs) specific to either CD8alpha or CD8beta to probe the mechanism of CD8alphabeta binding to pMHCI. The YTS156.7 mAb inhibits this interaction and blocks T cell activation. To elucidate the molecular basis for this inhibition, the crystal structure of the CD8alphabeta immunoglobulin-like ectodomains were determined in complex with mAb YTS156.7 Fab at 2.7 A resolution. The YTS156.7 epitope on CD8beta was identified and implies that residues in the CDR1 and CDR2-equivalent loops of CD8beta are occluded upon binding to class I pMHC. To further characterize the pMHCI/CD8alphabeta interaction, binding of class I tetramers to CD8alphabeta on the surface of T cells was assessed in the presence of anti-CD8 mAbs. In contrast to YTS156.7, mAb YTS105.18, which is specific for CD8alpha, does not inhibit binding of CD8alphabeta to class I tetramers, indicating the YTS105.18 epitope is not occluded in the pMHCI/CD8alphabeta complex. Together, these data indicate a model for the pMHCI/CD8alphabeta interaction similar to that observed for CD8alphaalpha in the CD8alphaalpha/pMHCI complex, but in which CD8alpha occupies the lower orientation (membrane proximal to the antigen presenting cell), and CD8beta occupies the upper position (membrane distal). The implication of this molecular assembly for the function of CD8alphabeta in T cell activation is discussed.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.