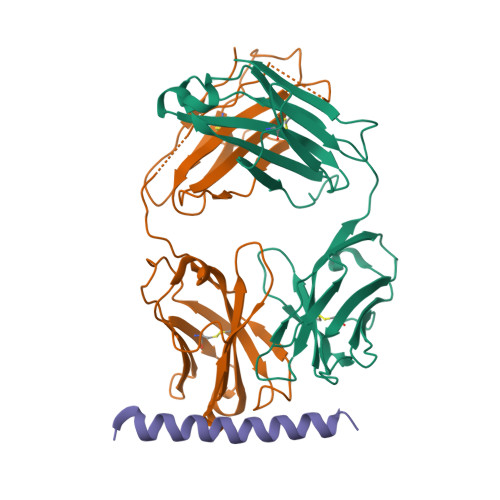
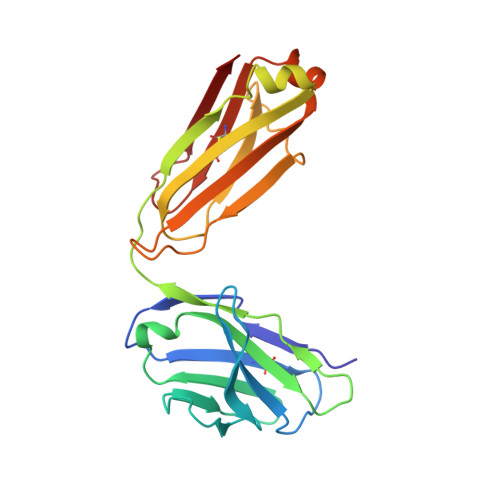
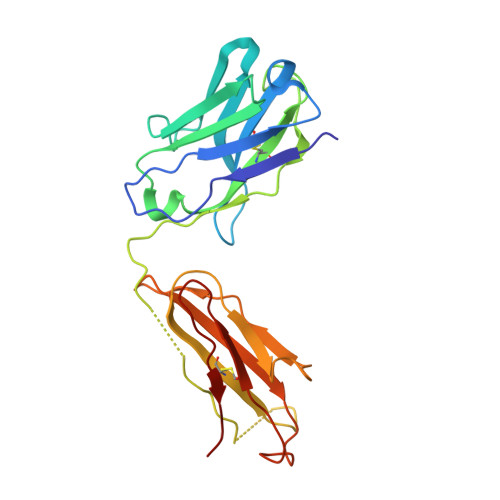
Crystal structure of full-length KcsA in its closed conformation.
Uysal, S., Vasquez, V., Tereshko, V., Esaki, K., Fellouse, F.A., Sidhu, S.S., Koide, S., Perozo, E., Kossiakoff, A.(2009) Proc Natl Acad Sci U S A 106: 6644-6649
- PubMed: 19346472
- DOI: https://doi.org/10.1073/pnas.0810663106
- Primary Citation of Related Structures:
3EFD, 3EFF - PubMed Abstract:
KcsA is a proton-activated, voltage-modulated K(+) channel that has served as the archetype pore domain in the Kv channel superfamily. Here, we have used synthetic antigen-binding fragments (Fabs) as crystallographic chaperones to determine the structure of full-length KcsA at 3.8 A, as well as that of its isolated C-terminal domain at 2.6 A. The structure of the full-length KcsA-Fab complex reveals a well-defined, 4-helix bundle that projects approximately 70 A toward the cytoplasm. This bundle promotes a approximately 15 degree bending in the inner bundle gate, tightening its diameter and shifting the narrowest point 2 turns of helix below. Functional analysis of the full-length KcsA-Fab complex suggests that the C-terminal bundle remains whole during gating. We suggest that this structure likely represents the physiologically relevant closed conformation of KcsA.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, and Institute for Biophysical Dynamics, University of Chicago, Chicago, IL 60637, USA.