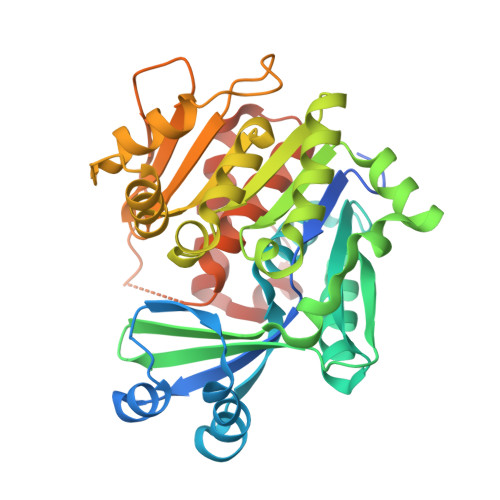
A High-Affinity Adenosine Kinase from Anopheles gambiae.
Cassera, M.B., Ho, M.C., Merino, E.F., Burgos, E.S., Rinaldo-Matthis, A., Almo, S.C., Schramm, V.L.(2011) Biochemistry 50: 1885-1893
- PubMed: 21247194
- DOI: https://doi.org/10.1021/bi101921w
- Primary Citation of Related Structures:
3LOO - PubMed Abstract:
Genome analysis revealed a mosquito orthologue of adenosine kinase in Anopheles gambiae (AgAK; the most important vector for the transmission of Plasmodium falciparum in Africa). P. falciparum are purine auxotrophs and do not express an adenosine kinase but rely on their hosts for purines. AgAK was kinetically characterized and found to have the highest affinity for adenosine (K(m) = 8.1 nM) of any known adenosine kinase. AgAK is specific for adenosine at the nucleoside site, but several nucleotide triphosphate phosphoryl donors are tolerated. The AgAK crystal structure with a bound bisubstrate analogue Ap(4)A (2.0 Å resolution) reveals interactions for adenosine and ATP and the geometry for phosphoryl transfer. The polyphosphate charge is partly neutralized by a bound Mg(2+) ion and an ion pair to a catalytic site Arg. The AgAK structure consists of a large catalytic core in a three-layer α/β/α sandwich, and a small cap domain in contact with adenosine. The specificity and tight binding for adenosine arise from hydrogen bond interactions of Asn14, Leu16, Leu40, Leu133, Leu168, Phe168, and Thr171 and the backbone of Ile39 and Phe168 with the adenine ring as well as through hydrogen bond interactions between Asp18, Gly64, and Asn68 and the ribosyl 2'- and 3'-hydroxyl groups. The structure is more similar to that of human adenosine kinase (48% identical) than to that of AK from Toxoplasma gondii (31% identical). With this extraordinary affinity for AgAK, adenosine is efficiently captured and converted to AMP at near the diffusion limit, suggesting an important role for this enzyme in the maintenance of the adenine nucleotide pool. mRNA analysis verifies that AgAK transcripts are produced in the adult insects.
Organizational Affiliation:
Department of Biochemistry, Albert Einstein College of Medicine, Yeshiva University, Bronx, New York 10461, United States.