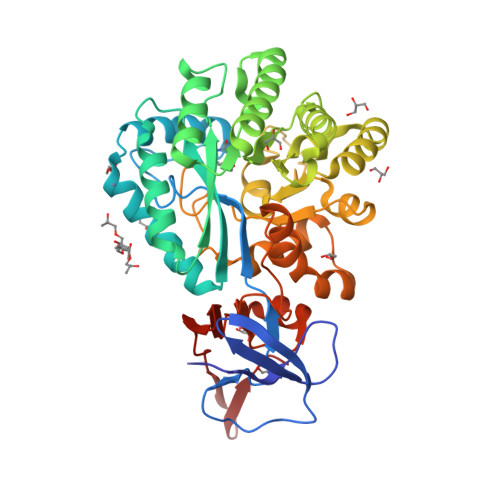
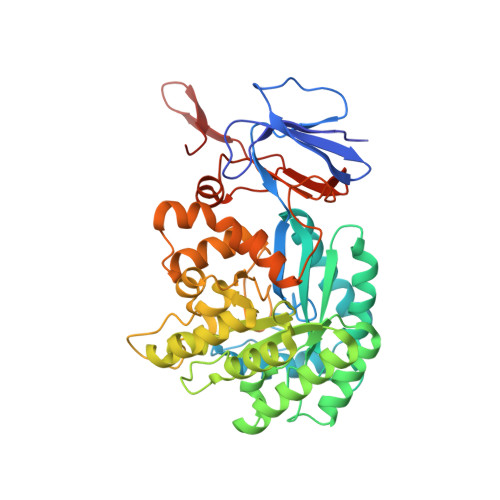
Three-dimensional structure and catalytic mechanism of Cytosine deaminase.
Hall, R.S., Fedorov, A.A., Xu, C., Fedorov, E.V., Almo, S.C., Raushel, F.M.(2011) Biochemistry 50: 5077-5085
- PubMed: 21545144
- DOI: https://doi.org/10.1021/bi200483k
- Primary Citation of Related Structures:
3O7U - PubMed Abstract:
Cytosine deaminase (CDA) from E. coli is a member of the amidohydrolase superfamily. The structure of the zinc-activated enzyme was determined in the presence of phosphonocytosine, a mimic of the tetrahedral reaction intermediate. This compound inhibits the deamination of cytosine with a K(i) of 52 nM. The zinc- and iron-containing enzymes were characterized to determine the effect of the divalent cations on activation of the hydrolytic water. Fe-CDA loses activity at low pH with a kinetic pK(a) of 6.0, and Zn-CDA has a kinetic pK(a) of 7.3. Mutation of Gln-156 decreased the catalytic activity by more than 5 orders of magnitude, supporting its role in substrate binding. Mutation of Glu-217, Asp-313, and His-246 significantly decreased catalytic activity supporting the role of these three residues in activation of the hydrolytic water molecule and facilitation of proton transfer reactions. A library of potential substrates was used to probe the structural determinants responsible for catalytic activity. CDA was able to catalyze the deamination of isocytosine and the hydrolysis of 3-oxauracil. Large inverse solvent isotope effects were obtained on k(cat) and k(cat)/K(m), consistent with the formation of a low-barrier hydrogen bond during the conversion of cytosine to uracil. A chemical mechanism for substrate deamination by CDA was proposed.
Organizational Affiliation:
Department of Chemistry, Texas A&M University, College Station, Texas 77842-3012, United States.