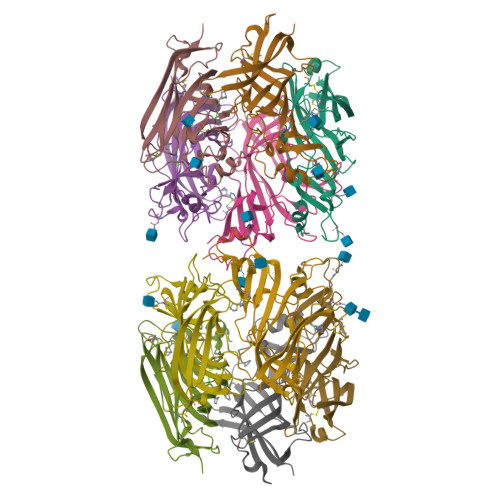
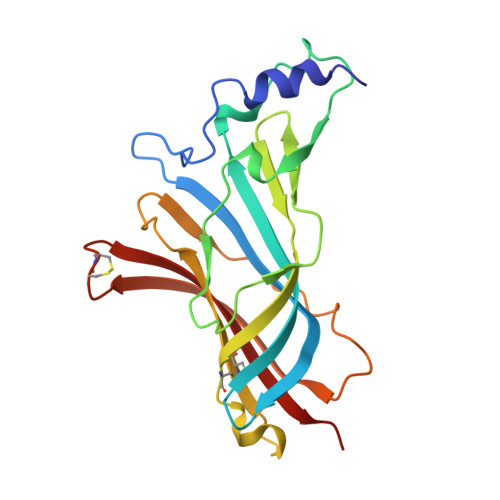
Ligand-binding domain of an alpha 7-nicotinic receptor chimera and its complex with agonist.
Li, S.X., Huang, S., Bren, N., Noridomi, K., Dellisanti, C.D., Sine, S.M., Chen, L.(2011) Nat Neurosci 14: 1253-1259
- PubMed: 21909087
- DOI: https://doi.org/10.1038/nn.2908
- Primary Citation of Related Structures:
3SQ6, 3SQ9 - PubMed Abstract:
The α(7) acetylcholine receptor (AChR) mediates pre- and postsynaptic neurotransmission in the central nervous system and is a potential therapeutic target in neurodegenerative, neuropsychiatric and inflammatory disorders. We determined the crystal structure of the extracellular domain of a receptor chimera constructed from the human α(7) AChR and Lymnaea stagnalis acetylcholine binding protein (AChBP), which shares 64% sequence identity and 71% similarity with native α(7). We also determined the structure with bound epibatidine, a potent AChR agonist. Comparison of the structures revealed molecular rearrangements and interactions that mediate agonist recognition and early steps in signal transduction in α(7) AChRs. The structures further revealed a ring of negative charge within the central vestibule, poised to contribute to cation selectivity. Structure-guided mutational studies disclosed distinctive contributions to agonist recognition and signal transduction in α(7) AChRs. The structures provide a realistic template for structure-aided drug design and for defining structure-function relationships of α(7) AChRs.
Organizational Affiliation:
Molecular and Computational Biology, Department of Biological Sciences, University of Southern California, Los Angeles, California, USA.