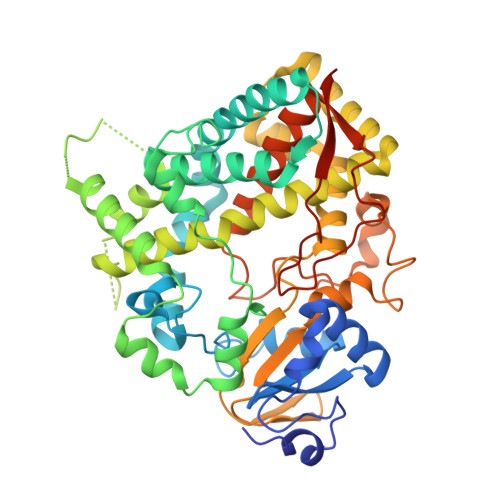
Structural and Mechanistic Insights into the Interaction of Cytochrome P4503A4 with Bromoergocryptine, a Type I Ligand.
Sevrioukova, I.F., Poulos, T.L.(2012) J Biol Chem 287: 3510-3517
- PubMed: 22157006
- DOI: https://doi.org/10.1074/jbc.M111.317081
- Primary Citation of Related Structures:
3UA1 - PubMed Abstract:
Cytochrome P4503A4 (CYP3A4), a major human drug-metabolizing enzyme, is responsible for the oxidation and clearance of the majority of administered drugs. One of the CYP3A4 substrates is bromoergocryptine (BEC), a dopamine receptor agonist prescribed for the inhibition of prolactin secretion and treatment of Parkinson disease, type 2 diabetes, and several other pathological conditions. Here we present a 2.15 Å crystal structure of the CYP3A4-BEC complex in which the drug, a type I heme ligand, is bound in a productive mode. The manner of BEC binding is consistent with the in vivo metabolite analysis and identifies the 8' and 9' carbons of the proline ring as the primary sites of oxidation. The crystal structure predicts the importance of Arg(212) and Thr(224) for binding of the tripeptide and lysergic moieties of BEC, respectively, which we confirmed experimentally. Our data support a three-step BEC binding model according to which the drug binds first at a peripheral site without perturbing the heme spectrum and then translocates into the active site cavity, where formation of a hydrogen bond between Thr(224) and the N1 atom of the lysergic moiety is followed by a slower conformational readjustment of the tripeptide group modulated by Arg(212).
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, California 92697-3900, USA.