Structure and mechanism of action of the hydroxy-aryl-aldehyde class of IRE1 endoribonuclease inhibitors.
Sanches, M., Duffy, N.M., Talukdar, M., Thevakumaran, N., Chiovitti, D., Canny, M.D., Lee, K., Kurinov, I., Uehling, D., Al-Awar, R., Poda, G., Prakesch, M., Wilson, B., Tam, V., Schweitzer, C., Toro, A., Lucas, J.L., Vuga, D., Lehmann, L., Durocher, D., Zeng, Q., Patterson, J.B., Sicheri, F.(2014) Nat Commun 5: 4202-4202
- PubMed: 25164867
- DOI: https://doi.org/10.1038/ncomms5202
- Primary Citation of Related Structures:
4PL3, 4PL4, 4PL5 - PubMed Abstract:
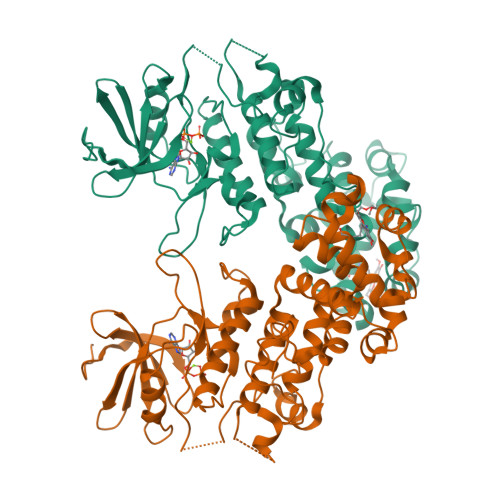
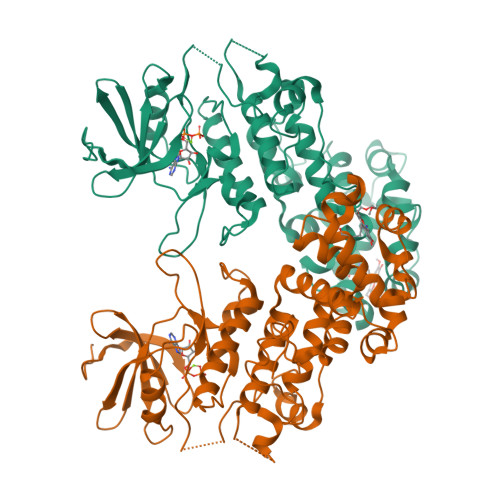
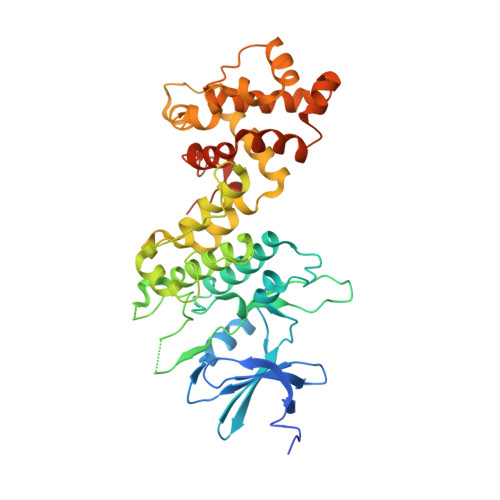
Endoplasmic reticulum (ER) stress activates the unfolded protein response and its dysfunction is linked to multiple diseases. The stress transducer IRE1α is a transmembrane kinase endoribonuclease (RNase) that cleaves mRNA substrates to re-establish ER homeostasis. Aromatic ring systems containing hydroxy-aldehyde moieties, termed hydroxy-aryl-aldehydes (HAA), selectively inhibit IRE1α RNase and thus represent a novel chemical series for therapeutic development. We solved crystal structures of murine IRE1α in complex with three HAA inhibitors. HAA inhibitors engage a shallow pocket at the RNase-active site through pi-stacking interactions with His910 and Phe889, an essential Schiff base with Lys907 and a hydrogen bond with Tyr892. Structure-activity studies and mutational analysis of contact residues define the optimal chemical space of inhibitors and validate the inhibitor-binding site. These studies lay the foundation for understanding both the biochemical and cellular functions of IRE1α using small molecule inhibitors and suggest new avenues for inhibitor design.
Organizational Affiliation:
1] Centre for Systems Biology, Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, Toronto, Ontario, Canada M5G 1X5 [2].