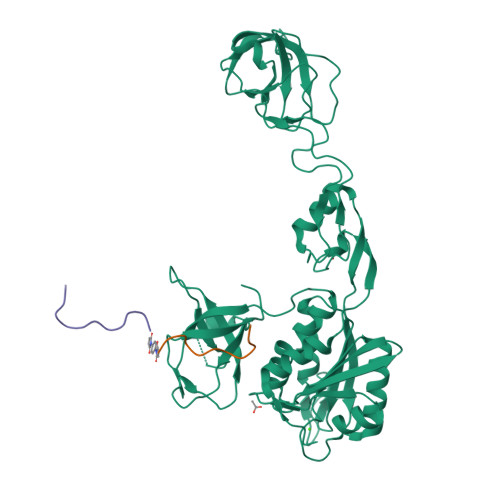
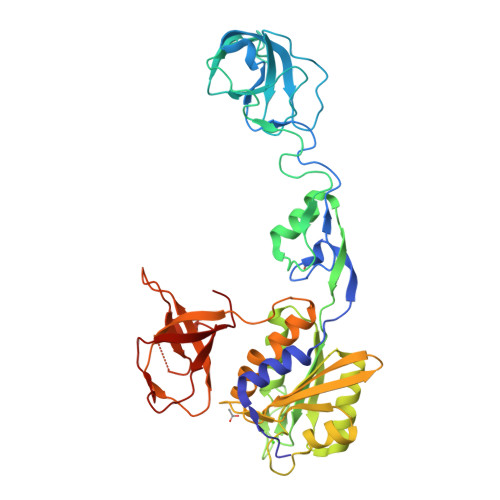
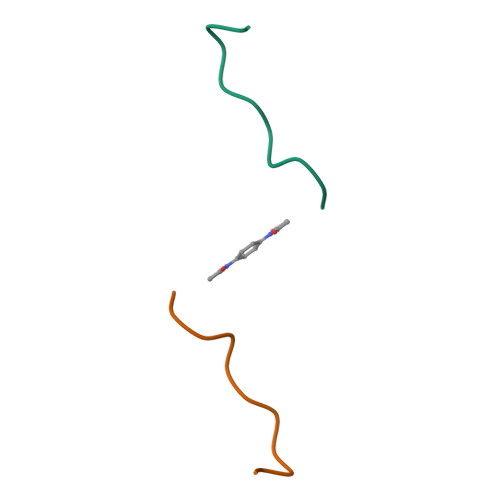
Design and Synthesis of High-Affinity Dimeric Inhibitors Targeting the Interactions between Gephyrin and Inhibitory Neurotransmitter Receptors.
Maric, H.M., Kasaragod, V.B., Haugaard-Kedstrom, L., Hausrat, T.J., Kneussel, M., Schindelin, H., Strmgaard, K.(2015) Angew Chem Int Ed Engl 54: 490-494
- PubMed: 25413248
- DOI: https://doi.org/10.1002/anie.201409043
- Primary Citation of Related Structures:
4U90, 4U91 - PubMed Abstract:
Gephyrin is the central scaffolding protein for inhibitory neurotransmitter receptors in the brain. Here we describe the development of dimeric peptides that inhibit the interaction between gephyrin and these receptors, a process which is fundamental to numerous synaptic functions and diseases of the brain. We first identified receptor-derived minimal gephyrin-binding peptides that displayed exclusive binding towards native gephyrin from brain lysates. We then designed and synthesized a series of dimeric ligands, which led to a remarkable 1220-fold enhancement of the gephyrin affinity (KD=6.8 nM). In X-ray crystal structures we visualized the simultaneous dimer-to-dimer binding in atomic detail, revealing compound-specific binding modes. Thus, we defined the molecular basis of the affinity-enhancing effect of multivalent gephyrin inhibitors and provide conceptually novel compounds with therapeutic potential, which will allow further elucidation of the gephyrin-receptor interplay.
Organizational Affiliation:
Department of Drug Design and Pharmacology, University of Copenhagen, Universitetsparken 2, 2100 Copenhagen (Denmark).