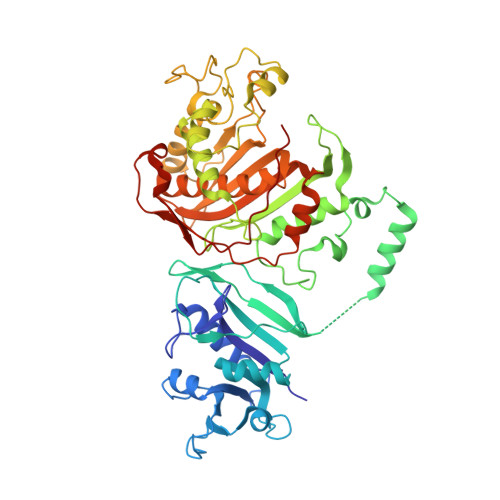
Substituted pyrrolo[2,3-d]pyrimidines as Cryptosporidium hominis thymidylate synthase inhibitors.
Kumar, V.P., Frey, K.M., Wang, Y., Jain, H.K., Gangjee, A., Anderson, K.S.(2013) Bioorg Med Chem Lett 23: 5426-5428
- PubMed: 23927969
- DOI: https://doi.org/10.1016/j.bmcl.2013.07.037
- Primary Citation of Related Structures:
4KY8 - PubMed Abstract:
Cryptosporidiosis, a gastrointestinal disease caused by a protozoan Cryptosporidium hominis is often fatal in immunocompromised individuals. There is little clinical data to show that the existing treatment by nitazoxanide and paromomycin is effective in immunocompromised individuals. Thymidylate synthase (TS) and dihydrofolate reductase (DHFR) are essential enzymes in the folate biosynthesis pathway and are well established as drug targets in cancer and malaria. A novel series of classical antifolates, 2-amino-4-oxo-5-substituted pyrrolo[2,3-d]pyrimidines have been evaluated as Cryptosporidium hominis thymidylate synthase (ChTS) inhibitors. Crystal structure in complex with the most potent compound, a 2'-chlorophenyl with a sulfur bridge with a Ki of 8.83±0.67 nM is discussed in terms of several Van der Waals, hydrophobic and hydrogen bond interactions with the protein residues and the substrate analog 5-fluorodeoxyuridine monophosphate. Of these interactions, two interactions with the non-conserved residues (A287 and S290) offer an opportunity to develop ChTS specific inhibitors. Compound 6 serves as a lead compound for analog design and its crystal structure provides clues for the design of ChTS specific inhibitors.
Organizational Affiliation:
Department of Pharmacology, Yale University School of Medicine, New Haven, CT 06520, USA.