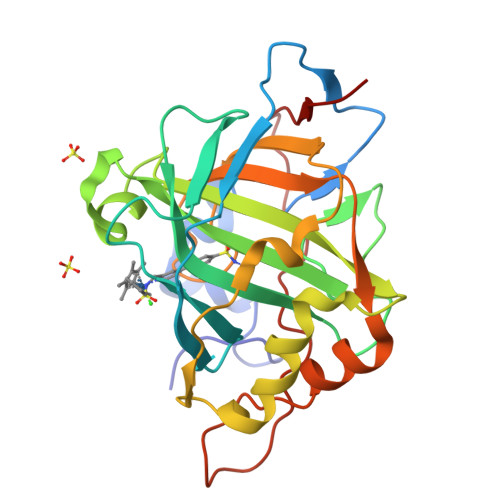
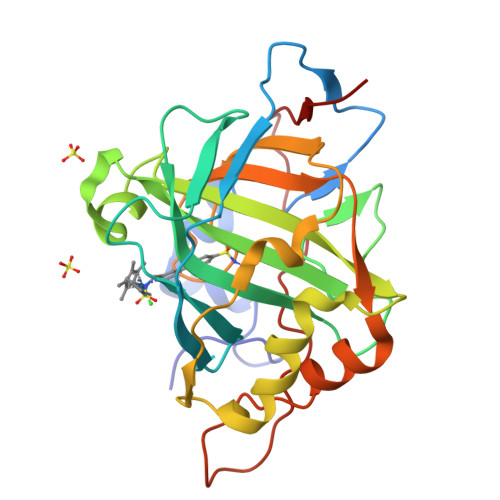
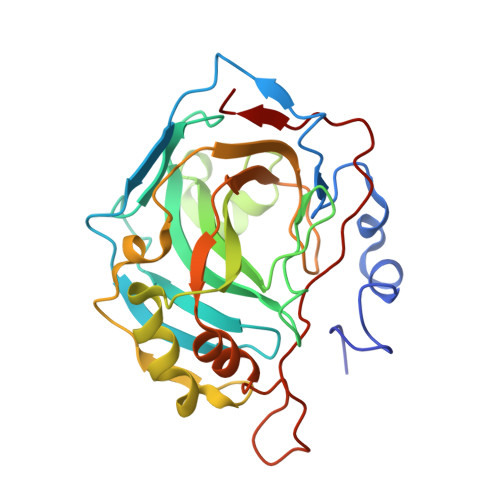
Improving the Catalytic Performance of an Artificial Metalloenzyme by Computational Design.
Heinisch, T., Pellizzoni, M., Durrenberger, M., Tinberg, C.E., Kohler, V., Klehr, J., Haussinger, D., Baker, D., Ward, T.R.(2015) J Am Chem Soc 137: 10414-10419
- PubMed: 26226626
- DOI: https://doi.org/10.1021/jacs.5b06622
- Primary Citation of Related Structures:
5BRU, 5BRV, 5BRW - PubMed Abstract:
Artifical metalloenzymes combine the reactivity of small molecule catalysts with the selectivity of enzymes, and new methods are required to tune the catalytic properties of these systems for an application of interest. Structure-based computational design could help to identify amino acid mutations leading to improved catalytic activity and enantioselectivity. Here we describe the application of Rosetta Design for the genetic optimization of an artificial transfer hydrogenase (ATHase hereafter), [(η(5)-Cp*)Ir(pico)Cl] ⊂ WT hCA II (Cp* = Me5C5(-)), for the asymmetric reduction of a cyclic imine, the precursor of salsolsidine. Based on a crystal structure of the ATHase, computational design afforded four hCAII variants with protein backbone-stabilizing and hydrophobic cofactor-embedding mutations. In dansylamide-competition assays, these designs showed 46-64-fold improved affinity for the iridium pianostool complex [(η(5)-Cp*)Ir(pico)Cl]. Gratifyingly, the new designs yielded a significant improvement in both activity and enantioselectivity (from 70% ee (WT hCA II) to up to 92% ee and a 4-fold increase in total turnover number) for the production of (S)-salsolidine. Introducing additional hydrophobicity in the Cp*-moiety of the Ir-catalyst provided by adding a propyl substituent on the Cp* moiety yields the most (S)-selective (96% ee) ATHase reported to date. X-ray structural data indicate that the high enantioselectivity results from embedding the piano stool moiety within the protein, consistent with the computational model.
Organizational Affiliation:
†Department of Chemistry, University of Basel, 4056 Basel, Switzerland.