Fragment-Linking Approach Using (19)F NMR Spectroscopy To Obtain Highly Potent and Selective Inhibitors of beta-Secretase.
Jordan, J.B., Whittington, D.A., Bartberger, M.D., Sickmier, E.A., Chen, K., Cheng, Y., Judd, T.(2016) J Med Chem 59: 3732-3749
- PubMed: 26978477
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01917
- Primary Citation of Related Structures:
5I3V, 5I3W, 5I3X, 5I3Y, 5IE1 - PubMed Abstract:
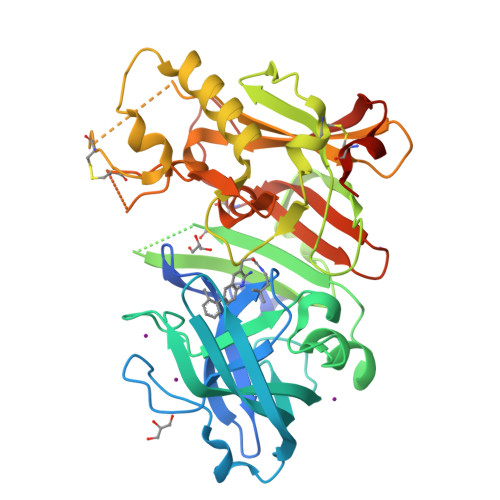
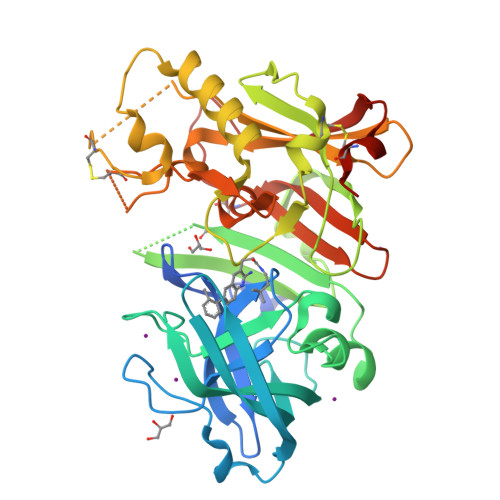
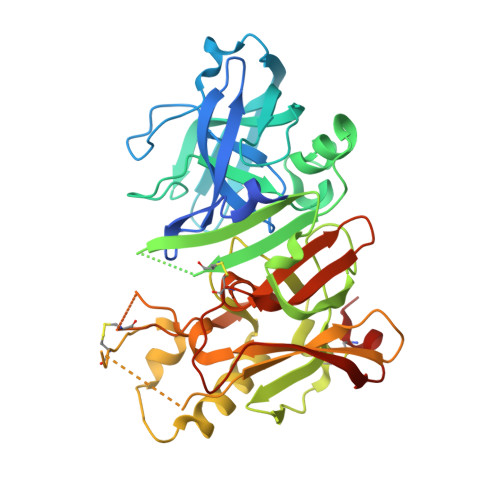
Fragment-based drug discovery (FBDD) has become a widely used tool in small-molecule drug discovery efforts. One of the most commonly used biophysical methods in detecting weak binding of fragments is nuclear magnetic resonance (NMR) spectroscopy. In particular, FBDD performed with (19)F NMR-based methods has been shown to provide several advantages over (1)H NMR using traditional magnetization-transfer and/or two-dimensional methods. Here, we demonstrate the utility and power of (19)F-based fragment screening by detailing the identification of a second-site fragment through (19)F NMR screening that binds to a specific pocket of the aspartic acid protease, β-secretase (BACE-1). The identification of this second-site fragment allowed the undertaking of a fragment-linking approach, which ultimately yielded a molecule exhibiting a more than 360-fold increase in potency while maintaining reasonable ligand efficiency and gaining much improved selectivity over cathepsin-D (CatD). X-ray crystallographic studies of the molecules demonstrated that the linked fragments exhibited binding modes consistent with those predicted from the targeted screening approach, through-space NMR data, and molecular modeling.
Organizational Affiliation:
Therapeutic Discovery, Amgen, Inc. One Amgen Center Drive, Thousand Oaks, California 91320, United States.