Comparative Analysis of Binding Kinetics and Thermodynamics of Dipeptidyl Peptidase-4 Inhibitors and Their Relationship to Structure.
Schnapp, G., Klein, T., Hoevels, Y., Bakker, R.A., Nar, H.(2016) J Med Chem 59: 7466-7477
- PubMed: 27438064
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00475
- Primary Citation of Related Structures:
5LLS - PubMed Abstract:
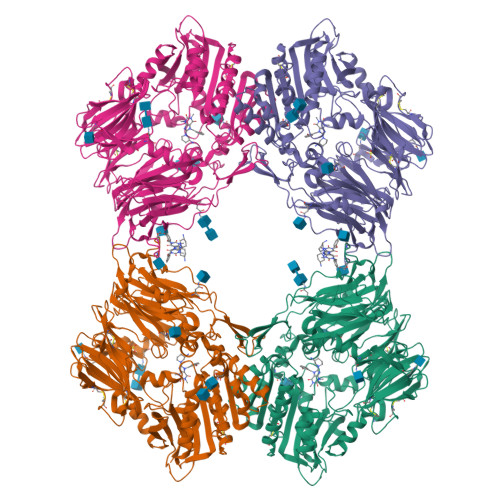
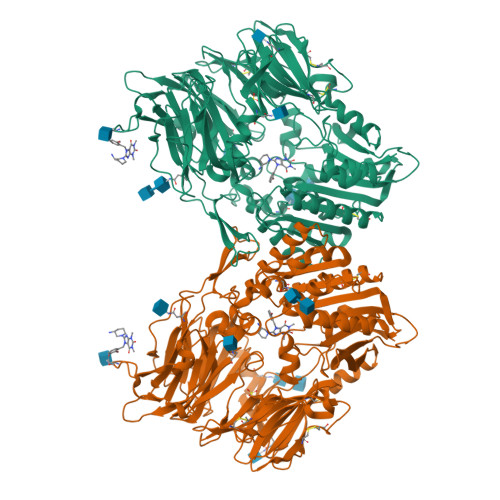
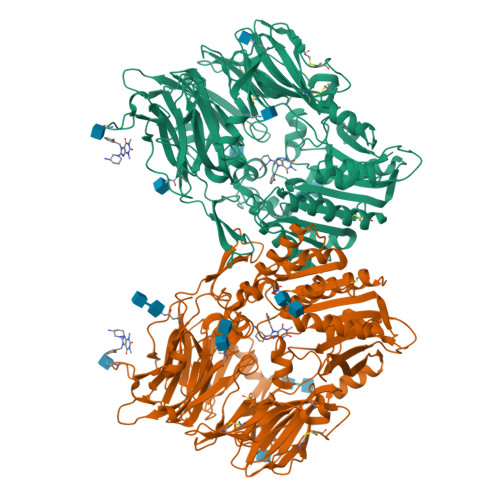
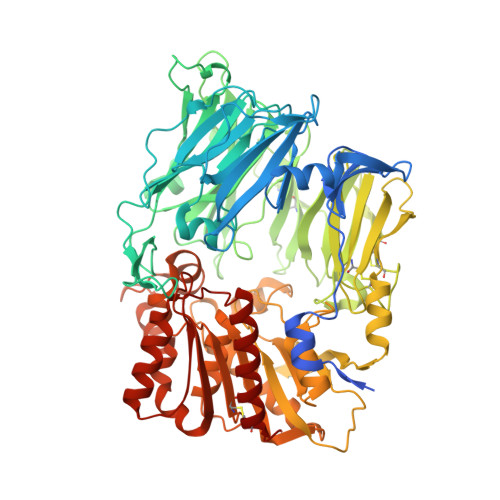
The binding kinetics and thermodynamics of dipeptidyl peptidase (DPP)-4 inhibitors (gliptins) were investigated using surface plasmon resonance and isothermal titration calorimetry. Binding of gliptins to DPP-4 is a rapid electrostatically driven process. Off-rates were generally slow partly because of reversible covalent bond formation by some gliptins, and partly because of strong and extensive interactions. Binding of all gliptins is enthalpy-dominated due to strong ionic interactions and strong solvent-shielded hydrogen bonds. Using a congeneric series of molecules which represented the intermediates in the lead optimization program of linagliptin, the onset of slow binding kinetics and development of the thermodynamic repertoire were analyzed in the context of incremental changes of the chemical structures. All compounds rapidly associated, and therefore the optimization of affinity and residence time is highly correlated. The major contributor to the increasing free energy of binding was a strong increase of binding enthalpy, whereas entropic contributions remained low and constant despite significant addition of lipophilicity.
Organizational Affiliation:
Department of Lead Identification and Optimization Support and ‡Department of CardioMetabolic Diseases Research, Boehringer Ingelheim Pharma GmbH & Co. KG , Biberach 88397, Germany.