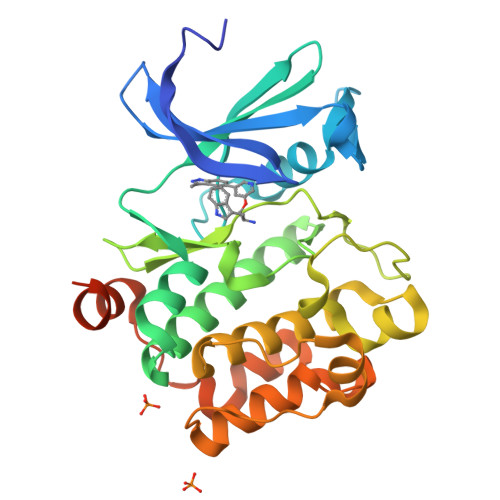
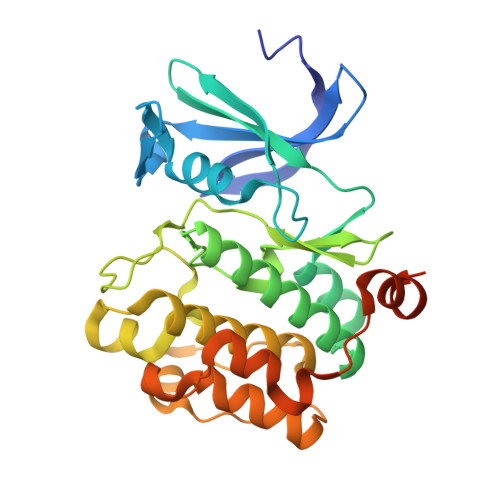
Discovery of 3,5-substituted 6-azaindazoles as potent pan-Pim inhibitors.
Hu, H., Wang, X., Chan, G.K., Chang, J.H., Do, S., Drummond, J., Ebens, A., Lee, W., Ly, J., Lyssikatos, J.P., Murray, J., Moffat, J.G., Chao, Q., Tsui, V., Wallweber, H., Kolesnikov, A.(2015) Bioorg Med Chem Lett 25: 5258-5264
- PubMed: 26459208
- DOI: https://doi.org/10.1016/j.bmcl.2015.09.052
- Primary Citation of Related Structures:
5DGZ, 5DHJ, 5DIA - PubMed Abstract:
Pim kinase inhibitors are promising cancer therapeutics. Pim-2, among the three Pim isoforms, plays a critical role in multiple myeloma yet inhibition of Pim-2 is challenging due to its high affinity for ATP. A co-crystal structure of a screening hit 1 bound to Pim-1 kinase revealed the key binding interactions of its indazole core within the ATP binding site. Screening of analogous core fragments afforded 1H-pyrazolo[3,4-c]pyridine (6-azaindazole) as a core for the development of pan-Pim inhibitors. Fragment and structure based drug design led to identification of the series with picomolar biochemical potency against all three Pim isoforms. Desirable cellular potency was also achieved.
Organizational Affiliation:
Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, United States.