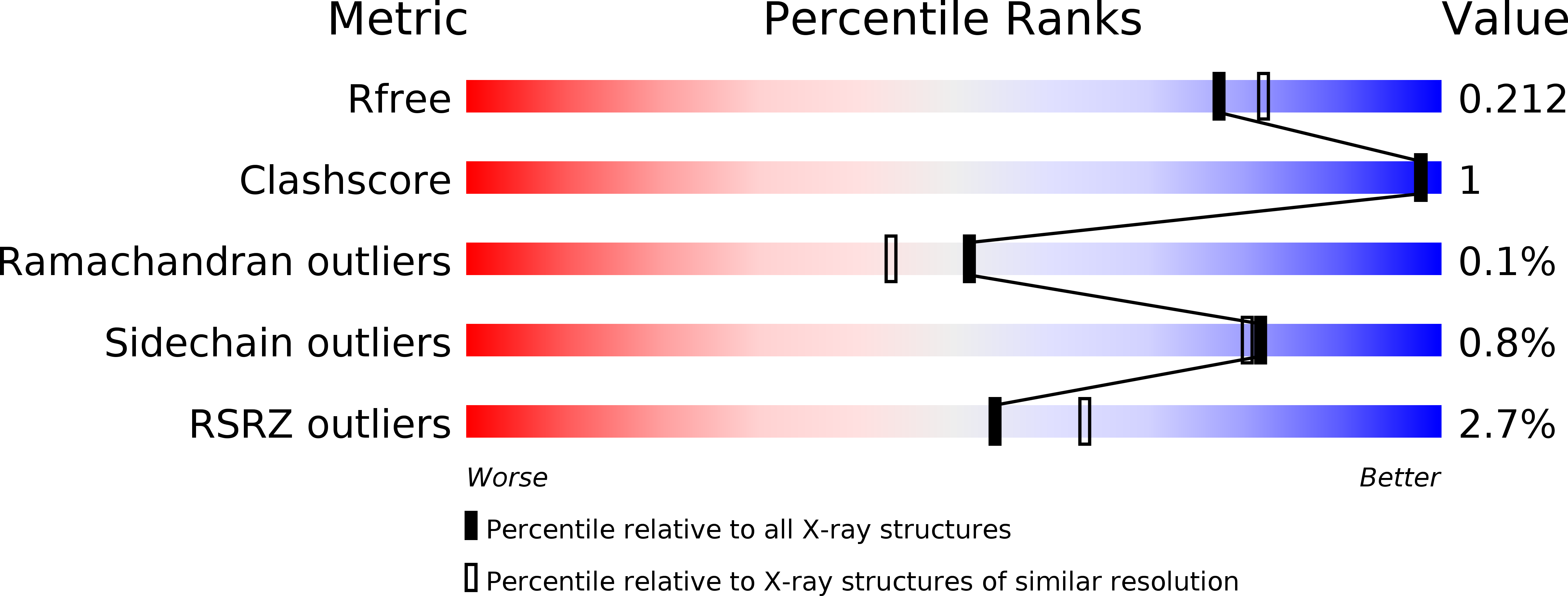
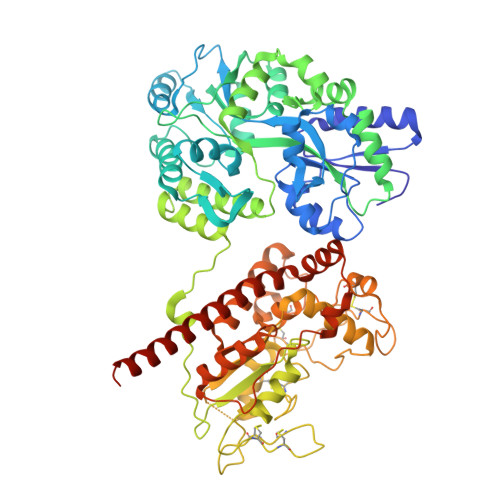
Structure Based Substrate Specificity Analysis of Heparan Sulfate 6-O-Sulfotransferases.
Xu, Y., Moon, A.F., Xu, S., Krahn, J.M., Liu, J., Pedersen, L.C.(2017) ACS Chem Biol 12: 73-82
- PubMed: 28103688
- DOI: https://doi.org/10.1021/acschembio.6b00841
- Primary Citation of Related Structures:
5T03, 5T05, 5T0A - PubMed Abstract:
Heparan sulfate (HS) is a sulfated polysaccharide exhibiting essential physiological functions. HS 6-O-sulfotransferase (6-OST) transfers a sulfo group to the 6-OH position of glucosamine units to confer a variety of HS biological activities. There are three different isoforms of 6-OST in the human genome. Here, we report crystal structures of the ternary complex of 6-OST with the sulfo donor analog 3'-phosphoadenosine 5'-phosphate and three different oligosaccharide substrates at 1.95 to 2.1 Å resolutions. Structural and mutational analyses reveal amino acid residues that contribute to catalysis and substrate recognition of 6-OST. Unexpectedly, the structures reveal 6-OST engages HS in a completely different orientation than other HS sulfotransferases and sheds light on the basic HS requirements for specificity. These findings also contribute structural information to understand mutations in human 6-OST isoform 1 associated with the human genetic disease idiopathic hypogonadotropic hypogonadism characterized by incomplete or lack of puberty.
Organizational Affiliation:
Division of Chemical Biology and Medicinal Chemistry, Eshelman School of Pharmacy, University of North Carolina , Chapel Hill, North Carolina 27599, United States.