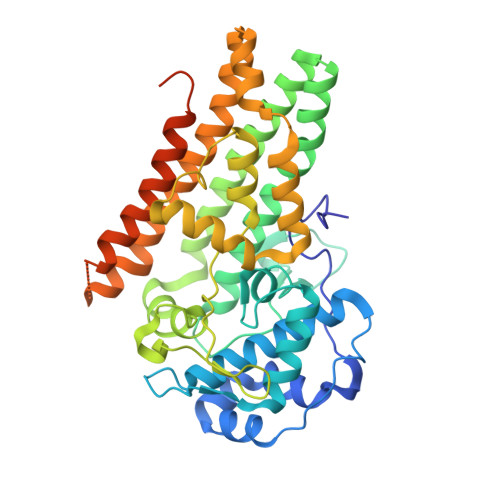
Structural insights into substrate and inhibitor binding sites in human indoleamine 2,3-dioxygenase 1.
Lewis-Ballester, A., Pham, K.N., Batabyal, D., Karkashon, S., Bonanno, J.B., Poulos, T.L., Yeh, S.R.(2017) Nat Commun 8: 1693-1693
- PubMed: 29167421
- DOI: https://doi.org/10.1038/s41467-017-01725-8
- Primary Citation of Related Structures:
5WMU, 5WMV, 5WMW, 5WMX, 5WN8 - PubMed Abstract:
Human indoleamine 2,3-dioxygenase 1 (hIDO1) is an attractive cancer immunotherapeutic target owing to its role in promoting tumoral immune escape. However, drug development has been hindered by limited structural information. Here, we report the crystal structures of hIDO1 in complex with its substrate, Trp, an inhibitor, epacadostat, and/or an effector, indole ethanol (IDE). The data reveal structural features of the active site (Sa) critical for substrate activation; in addition, they disclose a new inhibitor-binding mode and a distinct small molecule binding site (Si). Structure-guided mutation of a critical residue, F270, to glycine perturbs the Si site, allowing structural determination of an inhibitory complex, where both the Sa and Si sites are occupied by Trp. The Si site offers a novel target site for allosteric inhibitors and a molecular explanation for the previously baffling substrate-inhibition behavior of the enzyme. Taken together, the data open exciting new avenues for structure-based drug design.
Organizational Affiliation:
Department of Physiology and Biophysics, Albert Einstein College of Medicine, Bronx, NY, 10461, USA.