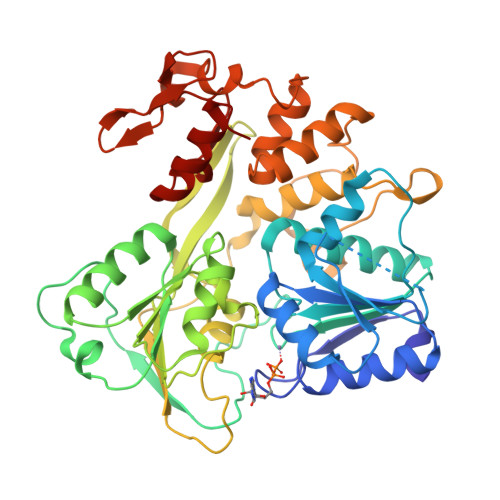
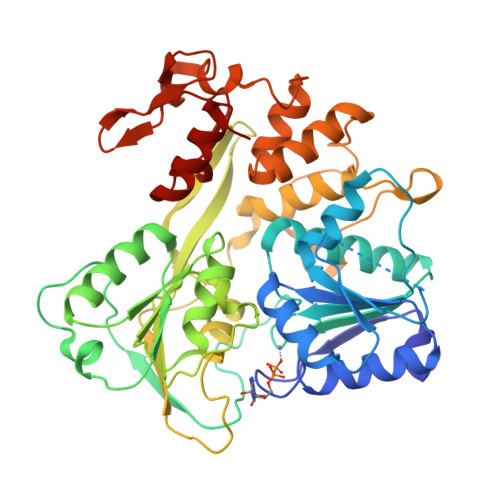
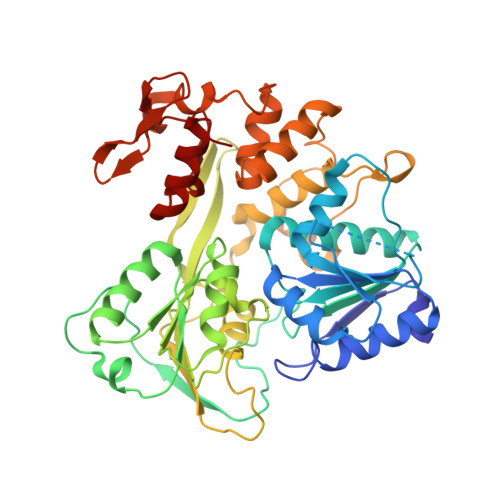
Mechanism of ATP hydrolysis by the Zika virus helicase.
Yang, X., Chen, C., Tian, H., Chi, H., Mu, Z., Zhang, T., Yang, K., Zhao, Q., Liu, X., Wang, Z., Ji, X., Yang, H.(2018) FASEB J 32: 5250-5257
- PubMed: 29913559
- DOI: https://doi.org/10.1096/fj.201701140R
- Primary Citation of Related Structures:
5Y6M, 5Y6N - PubMed Abstract:
During its life cycle, Zika virus (ZIKV), an arthropod-borne flavivirus that is associated with Guillain-Barré syndrome and causes microencephaly in fetuses and newborn children, encodes a critical and indispensable helicase domain that has 5'-triphosphatase activity and performs ATP hydrolysis to generate energy and thus, sustains unwinding of double-stranded RNA during ZIKV genome replication. Of these processes, ATP hydrolysis represents the most basic event; however, its dynamic mechanisms remain largely unknown, impeding the further understanding of the function of ZIKV helicase and the ongoing anti-ZIKV drug design. In this work, we determined the crystal structure of ZIKV helicase in complex with ADP-AlF 3 -Mn 2+ and ADP-Mn 2+ separately. The structural analysis indicates that these structures represent the intermediate state and posthydrolysis state, respectively, of the ATP hydrolysis process of ZIKV helicase. These findings, together with our earlier work, which identified the prehydrolysis state of ZIKV helicase, lead to a proposal of the ATP hydrolysis cycle for ZIKV helicase. On this basis, we used site-directed mutagenesis combined with an enzymatic study to identify successfully residues that are critical for the ATPase activity of ZIKV helicase; this will provide new ideas to understand the function for the key enzyme of ZIKV.-Yang, X., Chen, C., Tian, H., Chi, H., Mu, Z., Zhang, T., Yang, K., Zhao, Q., Liu, X., Wang, Z., Ji, X., Yang, H. Mechanism of ATP hydrolysis by the Zika virus helicase.
Organizational Affiliation:
School of Life Sciences, Tianjin University, Tianjin, China.