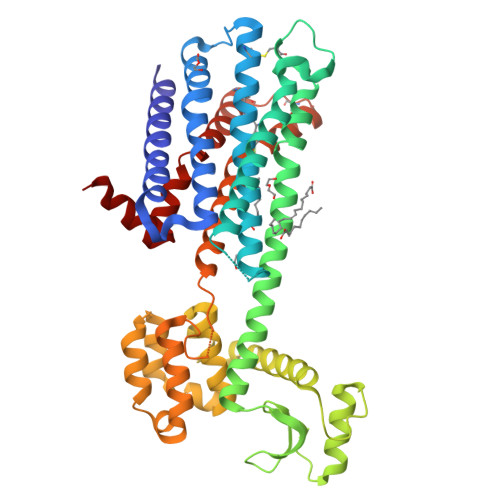
Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone.
Wang, S., Che, T., Levit, A., Shoichet, B.K., Wacker, D., Roth, B.L.(2018) Nature 555: 269-273
- PubMed: 29466326
- DOI: https://doi.org/10.1038/nature25758
- Primary Citation of Related Structures:
6CM4 - PubMed Abstract:
Dopamine is a neurotransmitter that has been implicated in processes as diverse as reward, addiction, control of coordinated movement, metabolism and hormonal secretion. Correspondingly, dysregulation of the dopaminergic system has been implicated in diseases such as schizophrenia, Parkinson's disease, depression, attention deficit hyperactivity disorder, and nausea and vomiting. The actions of dopamine are mediated by a family of five G-protein-coupled receptors. The D2 dopamine receptor (DRD2) is the primary target for both typical and atypical antipsychotic drugs, and for drugs used to treat Parkinson's disease. Unfortunately, many drugs that target DRD2 cause serious and potentially life-threatening side effects due to promiscuous activities against related receptors. Accordingly, a molecular understanding of the structure and function of DRD2 could provide a template for the design of safer and more effective medications. Here we report the crystal structure of DRD2 in complex with the widely prescribed atypical antipsychotic drug risperidone. The DRD2-risperidone structure reveals an unexpected mode of antipsychotic drug binding to dopamine receptors, and highlights structural determinants that are essential for the actions of risperidone and related drugs at DRD2.
Organizational Affiliation:
Department of Pharmacology, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599-7365, USA.