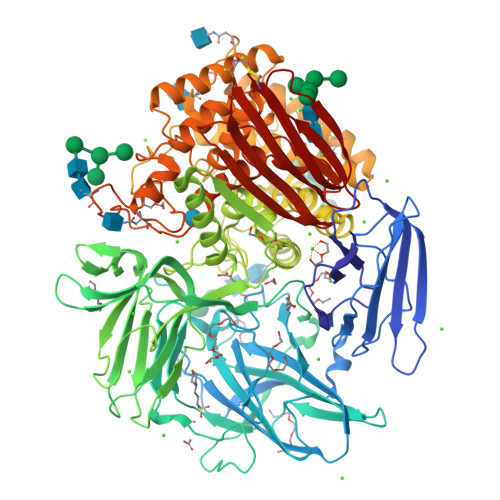
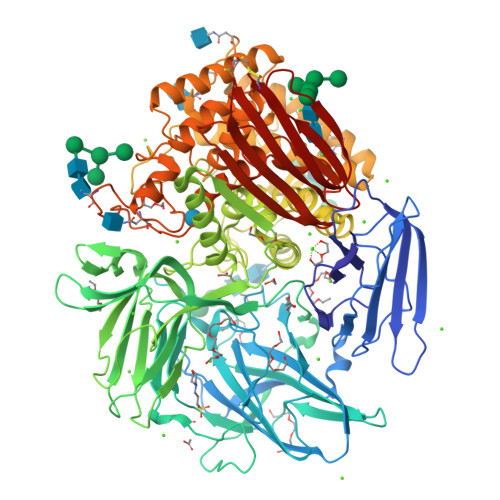
Crystal structure of native alpha-L-rhamnosidase from Aspergillus terreus.
Pachl, P., Skerlova, J., Simcikova, D., Kotik, M., Krenkova, A., Mader, P., Brynda, J., Kapesova, J., Kren, V., Otwinowski, Z., Rezacova, P.(2018) Acta Crystallogr D Struct Biol 74: 1078-1084
- PubMed: 30387766
- DOI: https://doi.org/10.1107/S2059798318013049
- Primary Citation of Related Structures:
6GSZ - PubMed Abstract:
α-L-Rhamnosidases cleave terminal nonreducing α-L-rhamnosyl residues from many natural rhamnoglycosides. This makes them catalysts of interest for various biotechnological applications. The X-ray structure of the GH78 family α-L-rhamnosidase from Aspergillus terreus has been determined at 1.38 Å resolution using the sulfur single-wavelength anomalous dispersion phasing method. The protein was isolated from its natural source in the native glycosylated form, and the active site contained a glucose molecule, probably from the growth medium. In addition to its catalytic domain, the α-L-rhamnosidase from A. terreus contains four accessory domains of unknown function. The structural data suggest that two of these accessory domains, E and F, might play a role in stabilizing the aglycon portion of the bound substrate.
Organizational Affiliation:
Structural Biology, Institute of Organic Chemistry and Biochemistry, The Czech Academy of Sciences, Flemingovo nám. 2, 16610 Prague 6, Czech Republic.